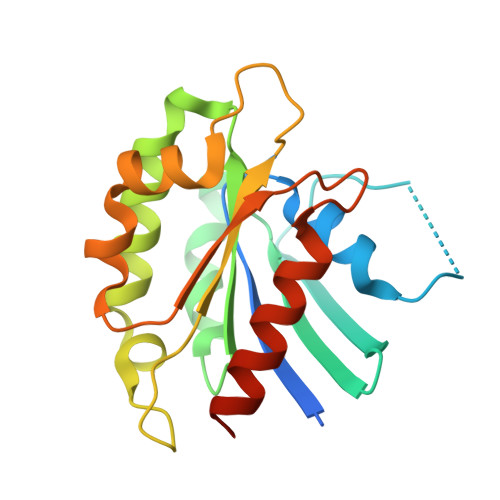
Crystal Structure of the Small Gtpase Arl6/Bbs3 from Trypanosomabrucei.
Hemsworth, G.R., Price, H.P., Smith, D.F., Wilson, K.S.(2013) Protein Sci 22: 196
- PubMed: 23184293 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2198
- Primary Citation Related Structures:
4BAS - PubMed Abstract:
Arl6/BBS3 is a small GTPase, mutations in which are implicated in the human ciliopathy Bardet-Biedl Syndrome (BBS). Arl6 is proposed to facilitate the recruitment of a large protein complex known as the BBSome to the base of the primary cilium, mediating specific trafficking of molecules to this important sensory organelle. Orthologues of Arl6 and the BBSome core subunits have been identified in the genomes of trypanosomes. Flagellum function and motility are crucial to the survival of Trypanosoma brucei, the causative agent of human African sleeping sickness, in the human bloodstream stage of its lifecycle and so the function of the BBSome proteins in trypanosomes warrants further study. RNAi knockdown of T. brucei Arl6 (TbArl6) has recently been shown to result in shortening of the trypanosome flagellum. Here we present the crystal structure of TbArl6 with the bound non-hydrolysable GTP analog GppNp at 2.0 Å resolution and highlight important differences between the trypanosomal and human proteins. Analysis of the TbArl6 active site confirms that it lacks the key glutamine that activates the nucleophile during GTP hydrolysis in other small GTPases. Furthermore, the trypanosomal proteins are significantly shorter at their N-termini suggesting a different method of membrane insertion compared to humans. Finally, analysis of sequence conservation suggests two surface patches that may be important for protein-protein interactions. Our structural analysis thus provides the basis for future biochemical characterisation of this important family of small GTPases.
- Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York, United Kingdom.
Organizational Affiliation: