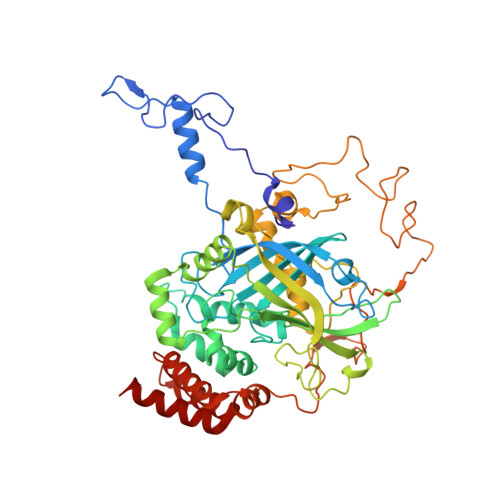
A Structural and Dynamic Investigation of the Inhibition of Catalase by Nitric Oxide.
Candelaresi, M., Gumiero, A., Adamczyk, K., Robb, K., Bellota-Anton, C., Sangal, V., Munnoch, J., Greetham, G.M., Towrie, M., Hoskisson, P.A., Parker, A.W., Tucker, N.P., Walsh, M.A., Hunt, N.T.(2013) Org Biomol Chem 11: 7778
- PubMed: 24121528 Search on PubMed
- DOI: https://doi.org/10.1039/c3ob41977k
- Primary Citation Related Structures:
4B7F, 4B7G, 4B7H - PubMed Abstract:
Determining the chemical and structural modifications occurring within a protein during fundamental processes such as ligand or substrate binding is essential to building up a complete picture of biological function. Currently, significant unanswered questions relate to the way in which protein structural dynamics fit within the structure-function relationship and to the functional role, if any, of bound water molecules in the active site. Addressing these questions requires a multidisciplinary approach and complementary experimental techniques that, in combination, enhance our understanding of the complexities of protein chemistry. We exemplify this philosophy by applying both physical and biological approaches to investigate the active site chemistry that contributes to the inhibition of the Corynebacterium glutamicum catalase enzyme by nitric oxide. Ultrafast two-dimensional infrared spectroscopy (2D-IR) experiments exploit the NO ligand as a local probe of the active site molecular environment and shows that catalase displays a dynamically-restricted, 'tight,' structure. X-ray crystallography studies of C. glutamicum catalase confirm the presence of a conserved chain of hydrogen-bonded bound water molecules that link the NO ligand and the protein scaffold. This combination of bound water and restricted dynamics stands in stark contrast to other haem proteins, such as myoglobin, that exhibit ligand transport functionality despite the presence of a similar distal architecture in close proximity to the ligand. We conclude not only that the bound water molecules in the catalase active site play an important role in molecular recognition of NO but also may be part of the mechanistic operation of this important enzyme.
- Department of Physics, University of Strathclyde, 107 Rottenrow East, Glasgow, G4 0NG, UK. neil.hunt@strath.ac.uk.
Organizational Affiliation: