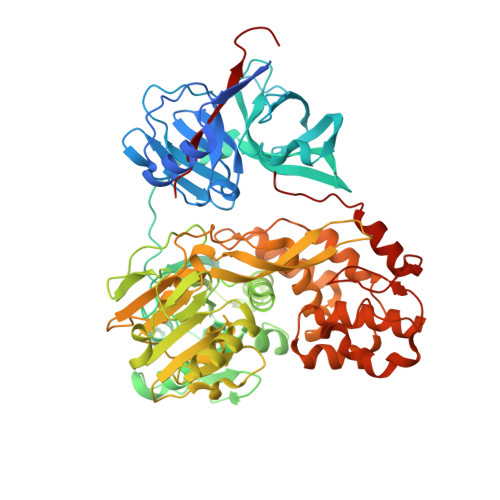
Discovery of an Allosteric Mechanism for the Regulation of Hcv Ns3 Protein Function.
Saalau-Bethell, S.M., Woodhead, A.J., Chessari, G., Carr, M.G., Coyle, J., Graham, B., Hiscock, S.D., Murray, C.W., Pathuri, P., Rich, S.J., Richardson, C.J., Williams, P.A., Jhoti, H.(2012) Nat Chem Biol 8: 920
- PubMed: 23023261 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1081
- Primary Citation Related Structures:
4B6E, 4B6F, 4B71, 4B73, 4B74, 4B75, 4B76 - PubMed Abstract:
Here we report a highly conserved new binding site located at the interface between the protease and helicase domains of the hepatitis C virus (HCV) NS3 protein. Using a chemical lead, identified by fragment screening and structure-guided design, we demonstrate that this site has a regulatory function on the protease activity via an allosteric mechanism. We propose that compounds binding at this allosteric site inhibit the function of the NS3 protein by stabilizing an inactive conformation and thus represent a new class of direct-acting antiviral agents.
- Astex Pharmaceuticals, Cambridge Science Park, Cambridge, UK.
Organizational Affiliation: