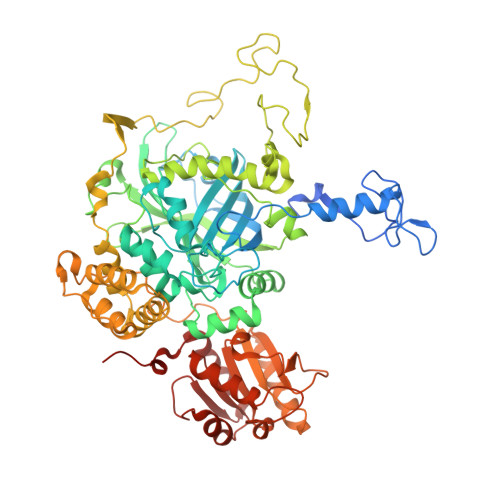
Investigating the Active Centre of the Scytalidium Thermophilum Catalase
Yuzugullu, Y., Trinh, C.H., Fairhurst, L., Ogel, Z.B., McPherson, M.J., Pearson, A.R.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 369
- PubMed: 23545640 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113004211
- Primary Citation Related Structures:
4B2Y, 4B31, 4B40, 4B5K - PubMed Abstract:
Almost all monofunctional haem catalases contain a highly conserved core containing the active site, which is connected to the exterior of the enzyme by three channels. These channels have been identified as potential routes for substrate flow and product release. To further investigate the role of these molecular channels, a series of mutants of Scytalidium thermophilum catalase were generated. The three-dimensional structures of four catalase variants, N155A, V123A, V123C and V123T, have been determined at resolutions of 2.25, 1.93, 1.9 and 1.7 Å, respectively. The V123C variant contains a new covalent bond between the S atom of Cys123 and the imidazole ring of the essential His82. This variant enzyme has only residual catalase activity and contains haem b instead of the normal haem d. The H82A variant demonstrates low catalase and phenol oxidase activities (0.2 and 20% of those of recombinant wild-type catalase-phenol oxidase, respectively). The N155A and N155H variants exhibit 4.5 and 3% of the wild-type catalase activity and contain haem d, showing that Asn155 is essential for catalysis but is not required for the conversion of haem b to haem d. Structural analysis suggests that the cause of the effect of these mutations on catalysis is the disruption of the ability of dioxygen substrates to efficiently access the active site. Additional mutants have been characterized biochemically to further probe the roles of the different channels. Introducing smaller or polar side chains in place of Val123 reduces the catalase activity. The F160V, F161V and F168V mutants show a marked decrease in catalase activity but have a much lower effect on the phenol oxidase activity, despite containing substoichiometric amounts of haem.
- Department of Biology, Kocaeli University, 41380 Kocaeli, Turkey.
Organizational Affiliation: