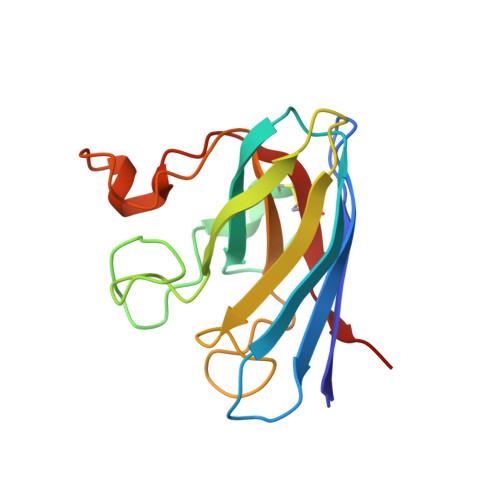
Structural Evidence for a Copper-Bound Carbonate Intermediate in the Peroxidase and Dismutase Activities of Superoxide Dismutase.
Strange, R.W., Hough, M.A., Antonyuk, S.V., Hasnain, S.S.(2012) PLoS One 7: 44811
- PubMed: 22984565
- DOI: https://doi.org/10.1371/journal.pone.0044811
- Primary Citation Related Structures:
4B3E - PubMed Abstract:
Copper-zinc superoxide dismutase (SOD) is of fundamental importance to our understanding of oxidative damage. Its primary function is catalysing the dismutation of superoxide to O(2) and H(2)O(2). SOD also reacts with H(2)O(2), leading to the formation of a strong copper-bound oxidant species that can either inactivate the enzyme or oxidise other substrates. In the presence of bicarbonate (or CO(2)) and H(2)O(2), this peroxidase activity is enhanced and produces the carbonate radical. This freely diffusible reactive oxygen species is proposed as the agent for oxidation of large substrates that are too bulky to enter the active site. Here, we provide direct structural evidence, from a 2.15 Å resolution crystal structure, of (bi)carbonate captured at the active site of reduced SOD, consistent with the view that a bound carbonate intermediate could be formed, producing a diffusible carbonate radical upon reoxidation of copper. The bound carbonate blocks direct access of substrates to Cu(I), suggesting that an adjunct to the accepted mechanism of SOD catalysed dismutation of superoxide operates, with Cu(I) oxidation by superoxide being driven via a proton-coupled electron transfer mechanism involving the bound carbonate rather than the solvent. Carbonate is captured in a different site when SOD is oxidised, being located in the active site channel adjacent to the catalytically important Arg143. This is the probable route of diffusion from the active site following reoxidation of the copper. In this position, the carbonate is poised for re-entry into the active site and binding to the reduced copper.
- Molecular Biophysics Group, Institute of Integrative Biology, Faculty of Health and Life Sciences, University of Liverpool, Liverpool, United Kingdom. r.strange@liv.ac.uk
Organizational Affiliation: