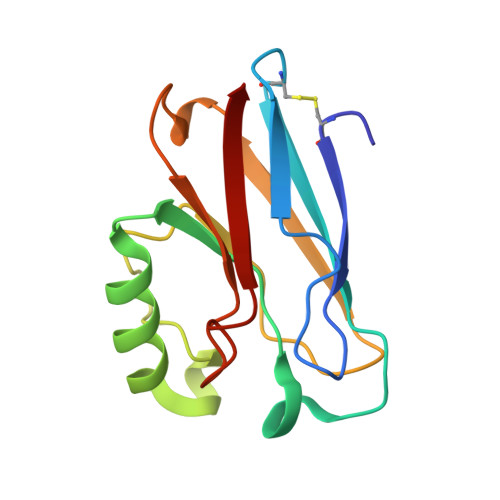
Crystal structure analysis of oxidized Pseudomonas aeruginosa azurin at pH 5.5 and pH 9.0. A pH-induced conformational transition involves a peptide bond flip.
Nar, H., Messerschmidt, A., Huber, R., van de Kamp, M., Canters, G.W.(1991) J Mol Biology 221: 765-772
- PubMed: 1942029 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)80173-r
- Primary Citation Related Structures:
4AZU, 5AZU - PubMed Abstract:
The X-ray crystal structure of recombinant wild-type azurin from Pseudomonas aeruginosa was determined by difference Fourier techniques using phases derived from the structure of the mutant His35Leu. Two data sets were collected from a single crystal of oxidized azurin soaked in mother liquor buffered at pH 5.5 and pH 9.0, respectively. Both data sets extend to 1.93 A resolution. The two pH forms were refined independently to crystallographic R-factors of 17.6% (pH 5.5) and 17.5% (pH 9.0). The conformational transition previously attributed to the protonation/deprotonation of residue His35 (pKa(red) = 7.3, pKa(ox) = 6.2), which lies in a crevice of the protein close to the copper binding site, involves a concomitant Pro36-Gly37 main-chain peptide bond flip. At the lower pH, the protonated imidazole N delta 1 of His35 forms a strong hydrogen bond with the carbonyl oxygen from Pro36, while at alkaline pH the deprotonated N delta 1 acts as an acceptor of a weak hydrogen bond from HN Gly37. The structure of the remainder of the azurin molecule, including the copper binding site, is not significantly affected by this transition.
- Max Planck Institut für Biochemie, Martinsried bei München, Germany.
Organizational Affiliation: