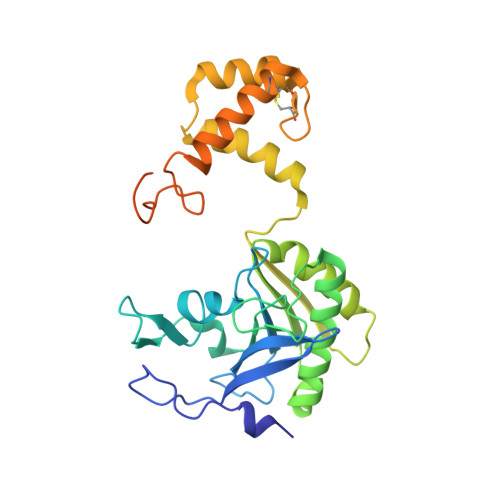
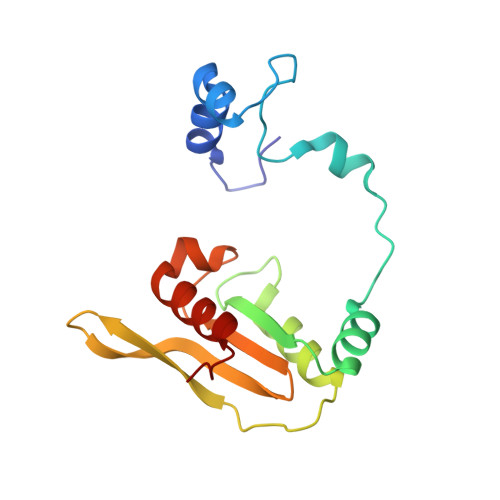
Novel Beta-Amino Acid Derivatives as Inhibitors of Cathepsin A.
Ruf, S., Buning, C., Schreuder, H., Horstick, G., Linz, W., Olpp, T., Pernerstorfer, J., Hiss, K., Kroll, K., Kannt, A., Kohlmann, M., Linz, D., Hubschle, T., Rutten, H., Wirth, K., Schmidt, T., Sadowski, T.(2012) J Med Chem 55: 7636
- PubMed: 22861813 Search on PubMed
- DOI: https://doi.org/10.1021/jm300663n
- Primary Citation Related Structures:
4AZ0, 4AZ3 - PubMed Abstract:
Cathepsin A (CatA) is a serine carboxypeptidase distributed between lysosomes, cell membrane, and extracellular space. Several peptide hormones including bradykinin and angiotensin I have been described as substrates. Therefore, the inhibition of CatA has the potential for beneficial effects in cardiovascular diseases. Pharmacological inhibition of CatA by the natural product ebelactone B increased renal bradykinin levels and prevented the development of salt-induced hypertension. However, so far no small molecule inhibitors of CatA with oral bioavailability have been described to allow further pharmacological profiling. In our work we identified novel β-amino acid derivatives as inhibitors of CatA after a HTS analysis based on a project adapted fragment approach. The new inhibitors showed beneficial ADME and pharmacokinetic profiles, and their binding modes were established by X-ray crystallography. Further investigations led to the identification of a hitherto unknown pathophysiological role of CatA in cardiac hypertrophy. One of our inhibitors is currently undergoing phase I clinical trials.
- Sanofi-Aventis Deutschland GmbH, Industriepark Höchst, 65926 Frankfurt, Germany. sven.ruf@sanofi.com
Organizational Affiliation: