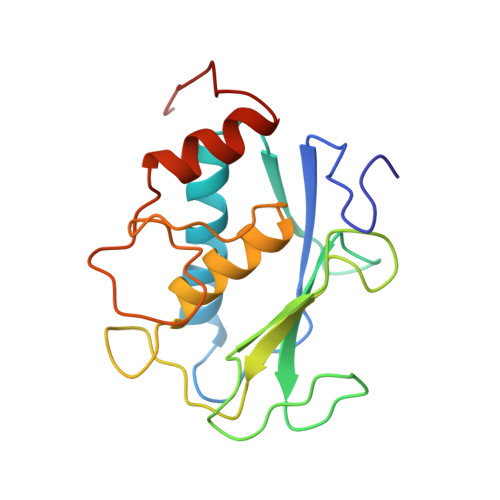
NMR solution structure of the catalytic fragment of human fibroblast collagenase complexed with a sulfonamide derivative of a hydroxamic acid compound.
Moy, F.J., Chanda, P.K., Chen, J.M., Cosmi, S., Edris, W., Skotnicki, J.S., Wilhelm, J., Powers, R.(1999) Biochemistry 38: 7085-7096
- PubMed: 10353819 Search on PubMed
- DOI: https://doi.org/10.1021/bi982576v
- Primary Citation Related Structures:
3AYK, 4AYK - PubMed Abstract:
The solution structure of the catalytic fragment of human fibroblast collagenase (MMP-1) complexed with a sulfonamide derivative of a hydroxamic acid compound (CGS-27023A) has been determined using two-dimensional and three-dimensional heteronuclear NMR spectroscopy. The solution structure of the complex was calculated by means of hybrid distance geometry-simulated annealing using a combination of experimental NMR restraints obtained from the previous refinement of the inhibitor-free MMP-1 (1) and recent restraints for the MMP-1:CGS-27023A complex. The hydroxamic acid moiety of CGS-27023A was found to chelate to the "right" of the catalytic zinc where the p-methoxyphenyl sits in the S1' active-site pocket, the isopropyl group is in contact with H83 and N80, and the pyridine ring is solvent exposed. The sulfonyl oxygens are in hydrogen-bonding distance to the backbone NHs of L81 and A82. This is similar to the conformation determined by NMR of the inhibitor bound to stromelysin (2, 3). A total of 48 distance restraints were observed between MMP-1 and CGS-27023A from 3D 13C-edited/12C-filtered NOESY and 3D 15N-edited NOESY experiments. An additional 18 intramolecular restraints were observed for CGS-27023A from a 2D 12C-filtered NOESY experiment. A minimal set of NMR experiments in combination with the free MMP-1 assignments were used to assign the MMP-1 (1)H, 13C, and 15N resonances in the MMP-1:CGS-27023A complex. The assignments of CGS-27023A in the complex were obtained from 2D 12C-filtered NOESY and 2D 12C-filtered TOCSY experiments.
- Department of Structural Biology, Wyeth-Ayerst Research, Pearl River, New York 10965, USA.
Organizational Affiliation: