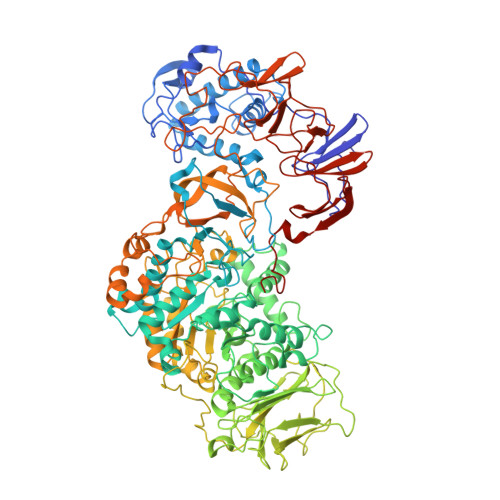
Flexibility of Truncated and Full-Length Glucansucrase Gtf180 Enzymes from Lactobacillus Reuteri 180.
Pijning, T., Vujicic-Zagar, A., Kralj, S., Dijkhuizen, L., Dijkstra, B.W.(2014) FEBS J 281: 2159
- PubMed: 24597929 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12769
- Primary Citation Related Structures:
4AYG - PubMed Abstract:
Glucansucrase enzymes synthesize high-molecular-mass extracellular α-glucan polysaccharides from sucrose. Previously, the crystal structure of truncated glucansucrase glucosyltransferase (GTF)180-ΔN from Lactobacillus reuteri 180 (lacking the N-terminal domain) revealed an elongated overall structure with two remote domains (IV and V) extending away from the core. By contrast, a new crystal form of the α-1,6/α-1,3 specific glucansucrase GTF180-ΔN shows an approximate 120(o) rotation of domain V about a hinge located between domains IV and V, giving a much more compact structure than before. Positional variability of domain V in solution is confirmed by small angle X-ray scattering experiments and rigid-body ensemble calculations. In addition, small angle X-ray scattering measurements of full-length GTF180 also provide the first structural data for a full-length glucansucrase, showing that the enzyme has an almost symmetric boomerang-like molecular shape, with a bend likely located between domains IV and V. The ~ 700-residue N-terminal domain, which is not present in the crystal structures, extends away from domain V and the catalytic core of the enzyme. We conclude that, as a result of the hinge region, in solution, GTF180-ΔN (and likely also the full-length GTF180) shows conformational flexibility; this may be a general feature of GH70 glucansucrases. • Structural data for GTF180-ΔN II have been deposited in the Protein Data Bank under accession code 4AYG.
- Laboratory of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute (GBB), University of Groningen, The Netherlands.
Organizational Affiliation: