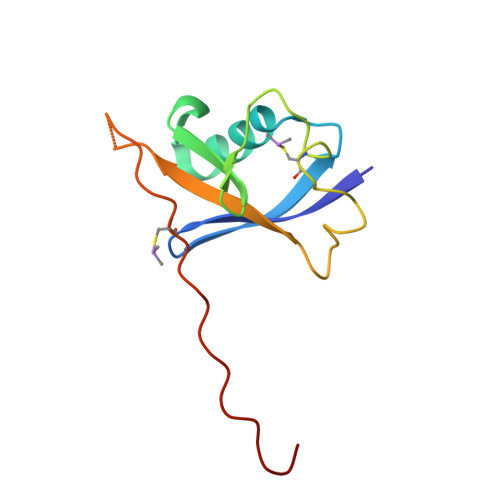
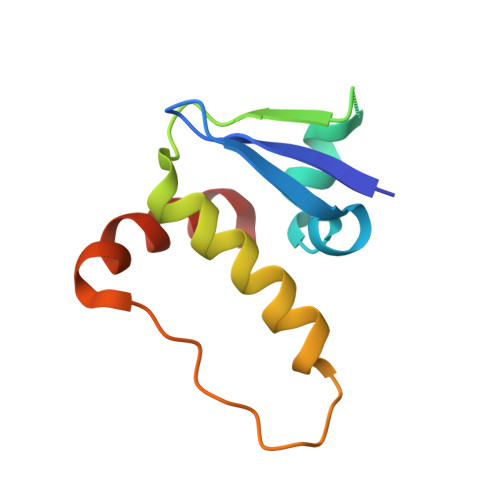
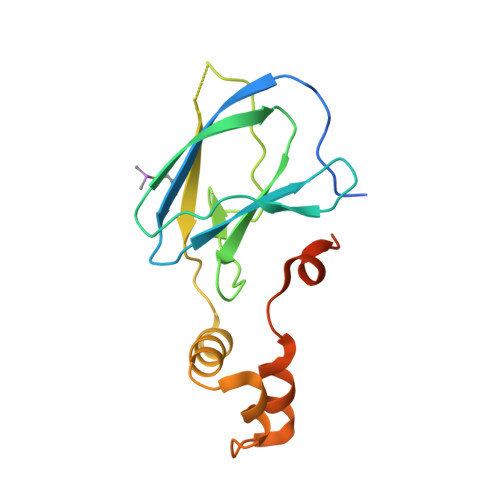
Dissecting Fragment-Based Lead Discovery at the Von Hippel-Lindau Protein:Hypoxia Inducible Factor 1Alpha Protein-Protein Interface.
Van Molle, I., Thomann, A., Buckley, D.L., So, E.C., Lang, S., Crews, C.M., Ciulli, A.(2012) Chem Biol 19: 1300
- PubMed: 23102223 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2012.08.015
- Primary Citation Related Structures:
3ZTC, 3ZTD, 4AJY, 4AWJ - PubMed Abstract:
Fragment screening is widely used to identify attractive starting points for drug design. However, its potential and limitations to assess the tractability of often challenging protein:protein interfaces have been underexplored. Here, we address this question by means of a systematic deconstruction of lead-like inhibitors of the pVHL:HIF-1α interaction into their component fragments. Using biophysical techniques commonly employed for screening, we could only detect binding of fragments that violate the Rule of Three, are more complex than those typically screened against classical druggable targets, and occupy two adjacent binding subsites at the interface rather than just one. Analyses based on ligand and group lipophilicity efficiency of anchored fragments were applied to dissect the individual subsites and probe for binding hot spots. The implications of our findings for targeting protein interfaces by fragment-based approaches are discussed.
- Department of Chemistry, University of Cambridge, Cambridge, CB2 1EW, UK.
Organizational Affiliation: