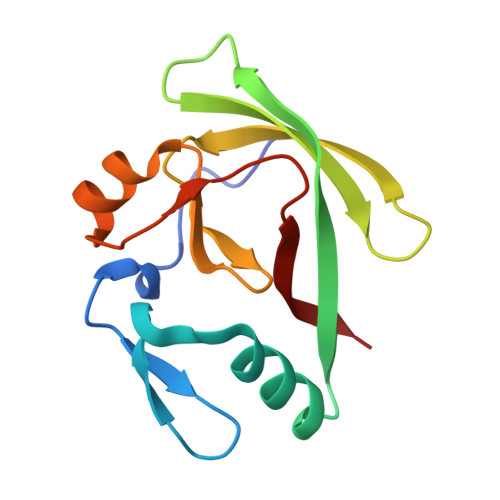
Self-Assembly and Conformational Heterogeneity of the Axh Domain of Ataxin-1: An Unusual Example of a Chameleon Fold
De Chiara, C., Rees, M., Menon, R.P., Pauwels, K., Lawrence, C., Konarev, P.V., Svergun, D.I., Martin, S.R., Chen, Y.W., Pastore, A.(2013) Biophys J 104: 1304
- PubMed: 23528090
- DOI: https://doi.org/10.1016/j.bpj.2013.01.048
- Primary Citation Related Structures:
4APT, 4AQP - PubMed Abstract:
Ataxin-1 is a human protein responsible for spinocerebellar ataxia type 1, a hereditary disease associated with protein aggregation and misfolding. Essential for ataxin-1 aggregation is the anomalous expansion of a polyglutamine tract near the protein N-terminus, but the sequence-wise distant AXH domain modulates and contributes to the process. The AXH domain is also involved in the nonpathologic functions of the protein, including a variety of intermolecular interactions with other cellular partners. The domain forms a globular dimer in solution and displays a dimer of dimers arrangement in the crystal asymmetric unit. Here, we have characterized the domain further by studying its behavior in the crystal and in solution. We solved two new structures of the domain crystallized under different conditions that confirm an inherent plasticity of the AXH fold. In solution, the domain is present as a complex equilibrium mixture of monomeric, dimeric, and higher molecular weight species. This behavior, together with the tendency of the AXH fold to be trapped in local conformations, and the multiplicity of protomer interfaces, makes the AXH domain an unusual example of a chameleon protein whose properties bear potential relevance for the aggregation properties of ataxin-1 and thus for disease.
- Medical Research Council National Institute for Medical Research London, United Kingdom.
Organizational Affiliation: