Functional and Structural Studies of a Novel Cold-Adapted Esterase from an Arctic Intertidal Metagenomic Library.
Fu, J., Leiros, H.-K.S., De Pascale, D., Johnson, K.A., Blencke, H.M., Landfald, B.(2013) Appl Microbiol Biotechnol 97: 3965
- PubMed: 22832985 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-012-4276-9
- Primary Citation Related Structures:
4AO6, 4AO7, 4AO8 - PubMed Abstract:
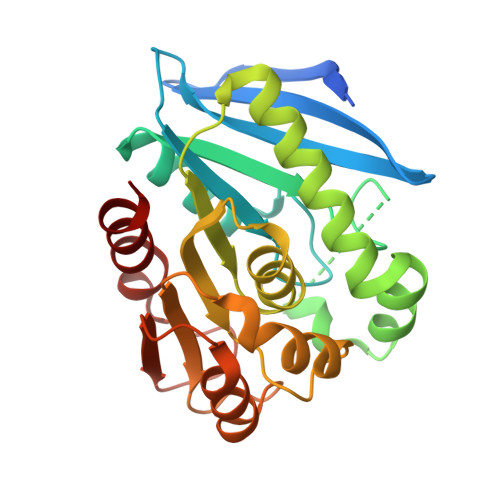
A novel cold-adapted lipolytic enzyme gene, est97, was identified from a high Arctic intertidal zone sediment metagenomic library. The deduced amino acid sequence of Est97 showed low similarity with other lipolytic enzymes, the maximum being 30 % identity with a putative lipase from Vibrio caribbenthicus. Common features of lipolytic enzymes, such as the GXSXG sequence motif, were detected. The gene product was over-expressed in Escherichia coli and purified. The recombinant Est97 (rEst97) hydrolysed various ρ-nitrophenyl esters with the best substrate being ρ-nitrophenyl hexanoate (K m and k cat of 39 μM and 25.8 s(-1), respectively). This esterase activity of rEst97 was optimal at 35 °C and pH 7.5 and the enzyme was unstable at temperatures above 25 °C. The apparent melting temperature, as determined by differential scanning calorimetry was 39 °C, substantiating Est97 as a cold-adapted esterase. The crystal structure of rEst97 was determined by the single wavelength anomalous dispersion method to 1.6 Å resolution. The protein was found to have a typical α/β-hydrolase fold with Ser144-His226-Asp197 as the catalytic triad. A suggested, relatively short lid domain of rEst97 is composed of residues 80-114, which form an α-helix and a disordered loop. The cold adaptation features seem primarily related to a high number of methionine and glycine residues and flexible loops in the high-resolution structures.
- Norwegian College of Fishery Science, University of Tromsø, 9037 Tromsø, Norway. juan.fu@uit.no
Organizational Affiliation: