Discovery and Optimization of New Benzimidazole- and Benzoxazole-Pyrimidone Selective Pi3Kbeta Inhibitors for the Treatment of Phosphatase and Tensin Homologue (Pten)-Deficient Cancers.
Certal, V., Halley, F., Virone-Oddos, A., Delorme, C., Karlsson, A., Rak, A., Thompson, F., Filoche-Romm, B., El-Ahmad, Y., Carry, J.C., Abecassis, P.Y., Lejeune, P., Vincent, L., Bonnevaux, H., Nicolas, J.P., Bertrand, T., Marquette, J.P., Michot, N., Benard, T., Below, P., Vade, I., Chatreaux, F., Lebourg, G., Pilorge, F., Angouillant-Boniface, O., Louboutin, A., Lengauer, C., Schio, L.(2012) J Med Chem 55: 4788
- PubMed: 22524426 Search on PubMed
- DOI: https://doi.org/10.1021/jm300241b
- Primary Citation Related Structures:
4AJW - PubMed Abstract:
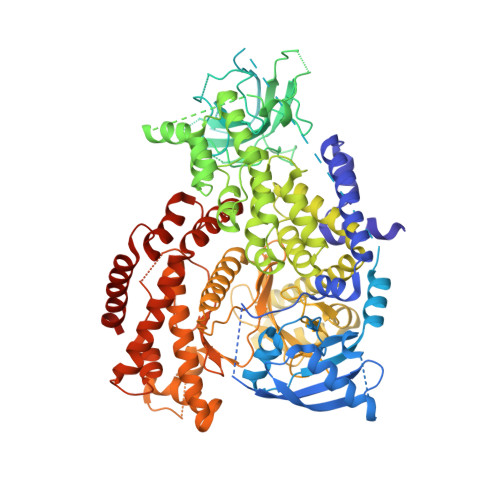
Most of the phosphoinositide-3 kinase (PI3K) kinase inhibitors currently in clinical trials for cancer treatment exhibit pan PI3K isoform profiles. Single PI3K isoforms differentially control tumorigenesis, and PI3Kβ has emerged as the isoform involved in the tumorigenicity of PTEN-deficient tumors. Herein we describe the discovery and optimization of a new series of benzimidazole- and benzoxazole-pyrimidones as small molecular mass PI3Kβ-selective inhibitors. Starting with compound 5 obtained from a one-pot reaction via a novel intermediate 1, medicinal chemistry optimization led to the discovery of compound 8, which showed a significant activity and selectivity for PI3Kβ and adequate in vitro pharmacokinetic properties. The X-ray costructure of compound 8 in PI3Kδ showed key interactions and structural features supporting the observed PI3Kβ isoform selectivity. Compound 8 achieved sustained target modulation and tumor growth delay at well tolerated doses when administered orally to SCID mice implanted with PTEN-deficient human tumor xenografts.
- Oncology Drug Discovery, Sanofi Research & Development , 13 quai Jules Guesde, 94403 Vitry-sur-Seine, France.
Organizational Affiliation: