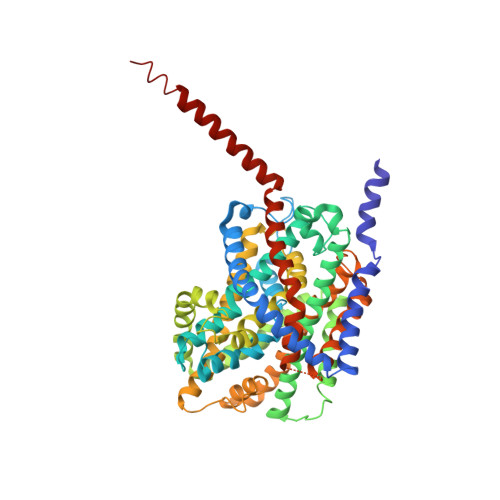
Alternating-Access Mechanism in Conformationally Asymmetric Trimers of the Betaine Transporter Betp.
Perez, C., Koshy, C., Yildiz, O., Ziegler, C.(2012) Nature 490: 126
- PubMed: 22940865 Search on PubMed
- DOI: https://doi.org/10.1038/nature11403
- Primary Citation Related Structures:
4AIN, 4DOJ - PubMed Abstract:
Betaine and Na(+) symport has been extensively studied in the osmotically regulated transporter BetP from Corynebacterium glutamicum, a member of the betaine/choline/carnitine transporter family, which shares the conserved LeuT-like fold of two inverted structural repeats. BetP adjusts its transport activity by sensing the cytoplasmic K(+) concentration as a measure for hyperosmotic stress via the osmosensing carboxy-terminal domain. BetP needs to be in a trimeric state for communication between individual protomers through several intratrimeric interaction sites. Recently, crystal structures of inward-facing BetP trimers have contributed to our understanding of activity regulation on a molecular level. Here we report new crystal structures, which reveal two conformationally asymmetric BetP trimers, capturing among them three distinct transport states. We observe a total of four new conformations at once: an outward-open apo and an outward-occluded apo state, and two closed transition states--one in complex with betaine and one substrate-free. On the basis of these new structures, we identified local and global conformational changes in BetP that underlie the molecular transport mechanism, which partially resemble structural changes observed in other sodium-coupled LeuT-like fold transporters, but show differences we attribute to the osmolytic nature of betaine, the exclusive substrate specificity and the regulatory properties of BetP.
- Max-Planck Institute of Biophysics, Department of Structural Biology, D-60438 Frankfurt am Main, Germany.
Organizational Affiliation: