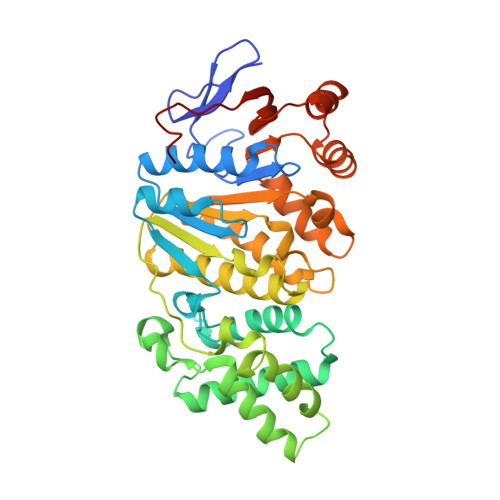
Structure of the Virb4 ATPase, Alone and Bound to the Core Complex of a Type Iv Secretion System.
Wallden, K., Williams, R., Yan, J., Lian, P.W., Wang, L., Thalassinos, K., Orlova, E.V., Waksman, G.(2012) Proc Natl Acad Sci U S A 109: 11348
- PubMed: 22745169 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1201428109
- Primary Citation Related Structures:
4AG5, 4AG6 - PubMed Abstract:
Type IV secretion (T4S) systems mediate the transfer of proteins and DNA across the cell envelope of bacteria. These systems play important roles in bacterial pathogenesis and in horizontal transfer of antibiotic resistance. The VirB4 ATPase of the T4S system is essential for both the assembly of the system and substrate transfer. In this article, we present the crystal structure of the C-terminal domain of Thermoanaerobacter pseudethanolicus VirB4. This structure is strikingly similar to that of another T4S ATPase, VirD4, a protein that shares only 12% sequence identity with VirB4. The VirB4 domain purifies as a monomer, but the full-length protein is observed in a monomer-dimer equilibrium, even in the presence of nucleotides and DNAs. We also report the negative stain electron microscopy structure of the core complex of the T4S system of the Escherichia coli pKM101 plasmid, with VirB4 bound. In this structure, VirB4 is also monomeric and bound through its N-terminal domain to the core's VirB9 protein. Remarkably, VirB4 is observed bound to the side of the complex where it is ideally placed to play its known regulatory role in substrate transfer.
- Institute of Structural and Molecular Biology, Department of Biological Sciences, Birkbeck, London WC1E 7HX, United Kingdom.
Organizational Affiliation: