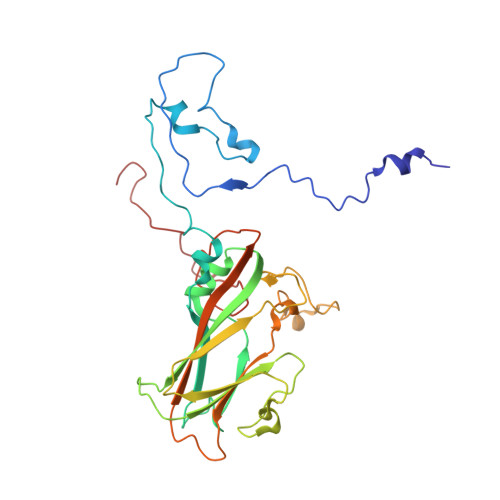
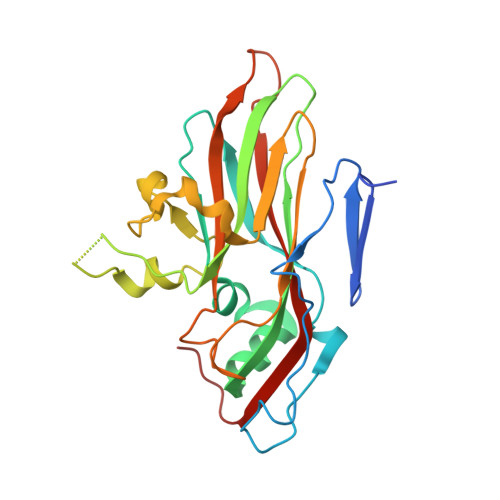
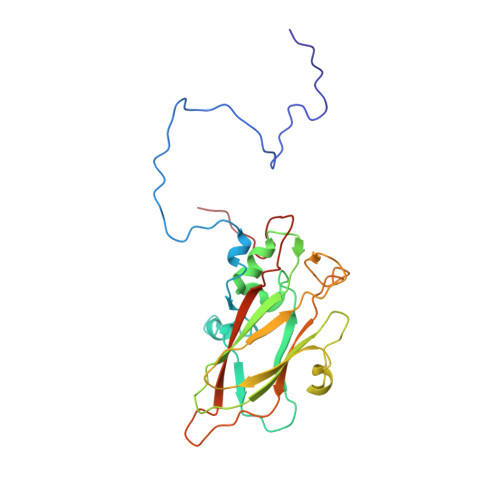
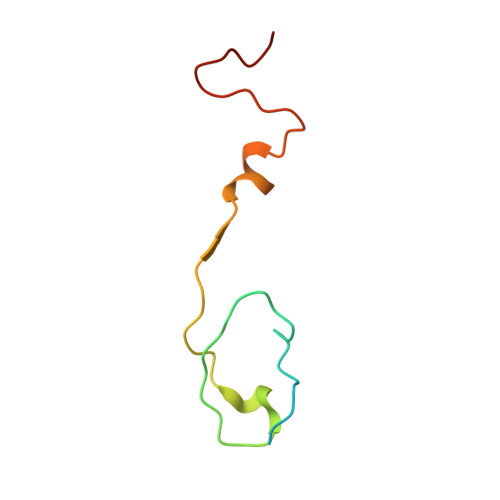
Crystal Structure of Human Enterovirus 71.
Plevka, P., Perera, R., Cardosa, J., Kuhn, R.J., Rossmann, M.G.(2012) Science 336: 1274
- PubMed: 22383808 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1218713
- Primary Citation Related Structures:
4AED - PubMed Abstract:
Enterovirus 71 is a picornavirus associated with fatal neurological illness in infants and young children. Here, we report the crystal structure of enterovirus 71 and show that, unlike in other enteroviruses, the "pocket factor," a small molecule that stabilizes the virus, is partly exposed on the floor of the "canyon." Thus, the structure of antiviral compounds may require a hydrophilic head group designed to interact with residues at the entrance of the pocket.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.
Organizational Affiliation: