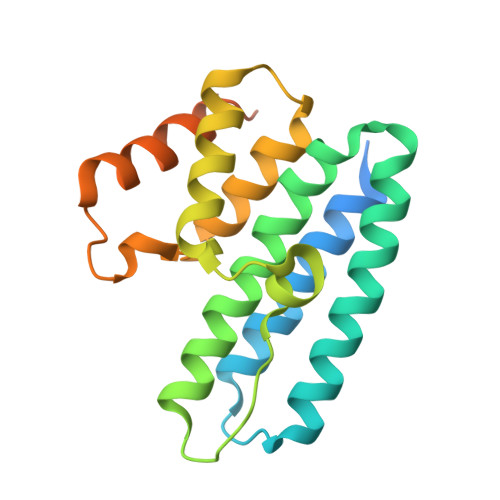
The Structure of the Conserved Type Six Secretion Protein Tssl (Dotu) from Francisella Novicida
Robb, C.S., Nano, F.E., Boraston, A.B.(2012) J Mol Biology 419: 277
- PubMed: 22504227 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.04.003
- Primary Citation Related Structures:
4ACK, 4ACL - PubMed Abstract:
Type six secretion systems (T6SSs) are found in many Gram-negative bacteria and are important for their virulence or their ecological competitiveness. The multicomponent T6SSs are responsible for the translocation of effector molecules into target eukaryotic or prokaryotic cells. The Francisella pathogenicity island encodes a putative T6SS that Francisella novicida requires for intramacrophage growth and virulence during infection of rodents. Here, we present the X-ray crystal structure of the conserved type six secretion component TssL (DotU) from F. novicida. The structure of this protein, which is referred to as Ftn_TssL, revealed an all-α-helical fold that is a unique fusion of two 3-helix bundles. The sequence of Ftn_TssL shows low identity to presumed homologs that are found in most T6SSs. The structure of Ftn_TssL, however, has allowed us to provide bioinformatics evidence that the F. novicida TssL has a fold that is very likely representative for TssL forms from both T6SSs and from the distantly related B subclass of type four secretion systems. A map of sequence conservation on the TssL structure revealed a surface-exposed groove that may represent a functional site on the protein.
- Department of Biochemistry and Microbiology, University of Victoria, PO Box 3055 Station CSC, Victoria, British Columbia, Canada V8W 3P6.
Organizational Affiliation: