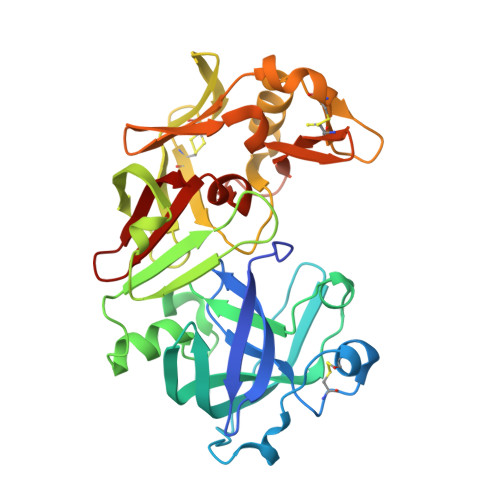
Camel and Bovine Chymosin: The Relationship between Their Structures and Cheese-Making Properties.
Langholm Jensen, J., Molgaard, A., Navarro Poulsen, J.C., Harboe, M.K., Simonsen, J.B., Lorentzen, A.M., Hjerno, K., van den Brink, J.M., Qvist, K.B., Larsen, S.(2013) Acta Crystallogr D Biol Crystallogr 69: 901-913
- PubMed: 23633601 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444913003260
- Primary Citation Related Structures:
4AA8, 4AA9 - PubMed Abstract:
Bovine and camel chymosin are aspartic peptidases that are used industrially in cheese production. They cleave the Phe105-Met106 bond of the milk protein κ-casein, releasing its predominantly negatively charged C-terminus, which leads to the separation of the milk into curds and whey. Despite having 85% sequence identity, camel chymosin shows a 70% higher milk-clotting activity than bovine chymosin towards bovine milk. The activities, structures, thermal stabilities and glycosylation patterns of bovine and camel chymosin obtained by fermentation in Aspergillus niger have been examined. Different variants of the enzymes were isolated by hydrophobic interaction chromatography and showed variations in their glycosylation, N-terminal sequences and activities. Glycosylation at Asn291 and the loss of the first three residues of camel chymosin significantly decreased its activity. Thermal differential scanning calorimetry revealed a slightly higher thermal stability of camel chymosin compared with bovine chymosin. The crystal structure of a doubly glycosylated variant of camel chymosin was determined at a resolution of 1.6 Å and the crystal structure of unglycosylated bovine chymosin was redetermined at a slightly higher resolution (1.8 Å) than previously determined structures. Camel and bovine chymosin share the same overall fold, except for the antiparallel central β-sheet that connects the N-terminal and C-terminal domains. In bovine chymosin the N-terminus forms one of the strands which is lacking in camel chymosin. This difference leads to an increase in the flexibility of the relative orientation of the two domains in the camel enzyme. Variations in the amino acids delineating the substrate-binding cleft suggest a greater flexibility in the ability to accommodate the substrate in camel chymosin. Both enzymes possess local positively charged patches on their surface that can play a role in interactions with the overall negatively charged C-terminus of κ-casein. Camel chymosin contains two additional positive patches that favour interaction with the substrate. The improved electrostatic interactions arising from variation in the surface charges and the greater malleability both in domain movements and substrate binding contribute to the better milk-clotting activity of camel chymosin towards bovine milk.
- Department of Chemistry, University of Copenhagen, Denmark.
Organizational Affiliation: