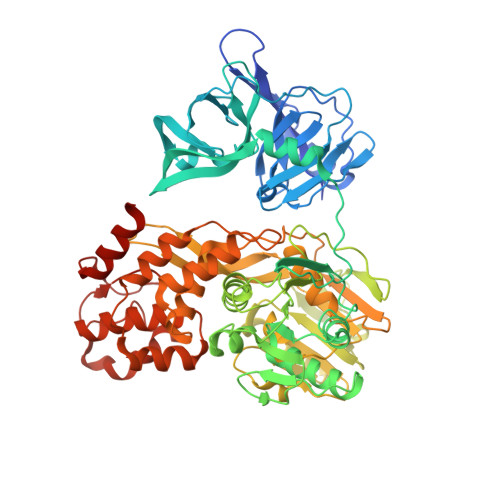
A Macrocyclic Hcv Ns3/4A Protease Inhibitor Interacts with Protease and Helicase Residues in the Complex with its Full- Length Target.
Schiering, N., D'Arcy, A., Villard, F., Simic, O., Kamke, M., Monnet, G., Hassiepen, U., Svergun, D.I., Pulfer, R., Eder, J., Raman, P., Bodendorf, U.(2011) Proc Natl Acad Sci U S A 108: 21052
- PubMed: 22160684 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1110534108
- Primary Citation Related Structures:
4A92 - PubMed Abstract:
Hepatitis C virus (HCV) infection is a global health burden with over 170 million people infected worldwide. In a significant portion of patients chronic hepatitis C infection leads to serious liver diseases, including fibrosis, cirrhosis, and hepatocellular carcinoma. The HCV NS3 protein is essential for viral polyprotein processing and RNA replication and hence viral replication. It is composed of an N-terminal serine protease domain and a C-terminal helicase/NTPase domain. For full activity, the protease requires the NS4A protein as a cofactor. HCV NS3/4A protease is a prime target for developing direct-acting antiviral agents. First-generation NS3/4A protease inhibitors have recently been introduced into clinical practice, markedly changing HCV treatment options. To date, crystal structures of HCV NS3/4A protease inhibitors have only been reported in complex with the protease domain alone. Here, we present a unique structure of an inhibitor bound to the full-length, bifunctional protease-helicase NS3/4A and show that parts of the P4 capping and P2 moieties of the inhibitor interact with both protease and helicase residues. The structure sheds light on inhibitor binding to the more physiologically relevant form of the enzyme and supports exploring inhibitor-helicase interactions in the design of the next generation of HCV NS3/4A protease inhibitors. In addition, small angle X-ray scattering confirmed the observed protease-helicase domain assembly in solution.
- Expertise Platform Proteases, Novartis Institutes for BioMedical Research, Fabrikstrasse 16, CH-4002 Basel, Switzerland. nikolaus.schiering@novartis.com
Organizational Affiliation: