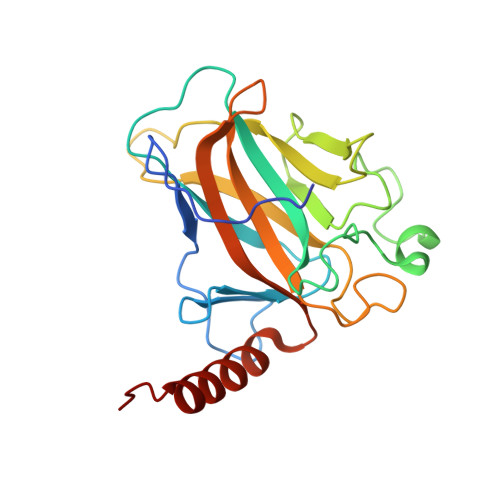
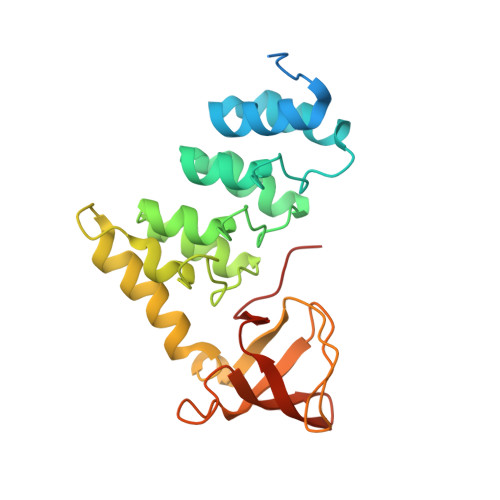
Structural Basis for Aspp2 Recognition by the Tumor Suppressor P73.
Canning, P., von Delft, F., Bullock, A.N.(2012) J Mol Biology 423: 515
- PubMed: 22917970 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2012.08.005
- Primary Citation Related Structures:
2XWC, 4A63 - PubMed Abstract:
Tumor suppressors p53, p63 and p73 comprise a family of stress-responsive transcription factors with distinct functions in development and tumor suppression. Most human cancers lose p53 function, yet all three proteins are capable of inducing apoptosis or cellular senescence. Mechanisms are therefore under investigation to activate p73-dependent apoptosis in p53-deficient cancer cells. Significantly, the DNA-binding domain (DBD) of p73 escapes viral oncoproteins and displays an enhanced thermal stability. To further understand the variant features of p73, we solved the high-resolution crystal structure of the p73 DBD as well as its complex with the ankyrin repeat and SH3 domains of the pro-apoptotic factor ASPP2. The p73 structure exhibits the same conserved architecture as p53 but displays a divergent L2 loop, a known site of protein-protein interaction. The loop in p73 is changed by a two-residue insertion that also induces repacking around the site of the p53 mutational hotspot R175. Importantly, the binding of ASPP2 is preserved by conformational changes in both the ankyrin repeat and SH3 domains. These results further highlight the structural variation that impacts p53 family interactions within the p53 interactome.
- Structural Genomics Consortium, University of Oxford, Old Road Campus, Roosevelt Drive, Oxford OX3 7DQ, UK.
Organizational Affiliation: