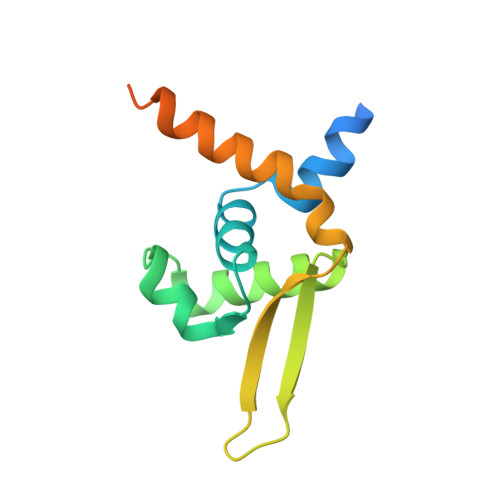
Structural Insights Into the Redox-Switch Mechanism of the Marr/Duf24-Type Regulator Hypr
Palm, G.J., Chi, B.K., Waack, P., Gronau, K., Becher, D., Albrecht, D., Hinrichs, W., Read, R.J., Antelmann, H.(2012) Nucleic Acids Res 40: 4178
- PubMed: 22238377 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkr1316
- Primary Citation Related Structures:
4A5M, 4A5N - PubMed Abstract:
Bacillus subtilis encodes redox-sensing MarR-type regulators of the OhrR and DUF24-families that sense organic hydroperoxides, diamide, quinones or aldehydes via thiol-based redox-switches. In this article, we characterize the novel redox-sensing MarR/DUF24-family regulator HypR (YybR) that is activated by disulphide stress caused by diamide and NaOCl in B. subtilis. HypR controls positively a flavin oxidoreductase HypO that confers protection against NaOCl stress. The conserved N-terminal Cys14 residue of HypR has a lower pK(a) of 6.36 and is essential for activation of hypO transcription by disulphide stress. HypR resembles a 2-Cys-type regulator that is activated by Cys14-Cys49' intersubunit disulphide formation. The crystal structures of reduced and oxidized HypR proteins were resolved revealing structural changes of HypR upon oxidation. In reduced HypR a hydrogen-bonding network stabilizes the reactive Cys14 thiolate that is 8-9 Å apart from Cys49'. HypR oxidation breaks these H-bonds, reorients the monomers and moves the major groove recognition α4 and α4' helices ∼4 Å towards each other. This is the first crystal structure of a redox-sensing MarR/DUF24 family protein in bacteria that is activated by NaOCl stress. Since hypochloric acid is released by activated macrophages, related HypR-like regulators could function to protect pathogens against the host immune defense.
- Institute for Biochemistry, Ernst-Moritz-Arndt-University of Greifswald, D-17487 Greifswald, Germany.
Organizational Affiliation: