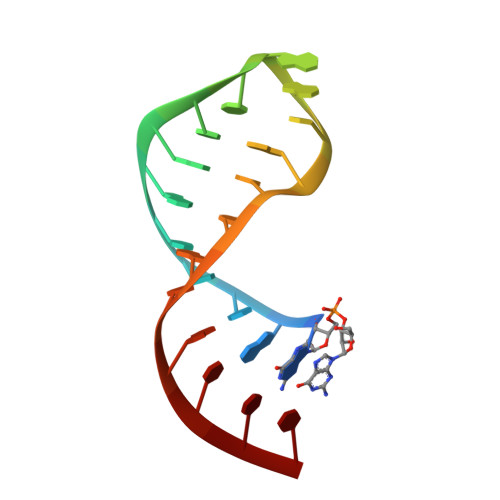
Unac Tetraloops: To What Extent Can They Mimic Gnra Tetraloops
Zhao, Q., Huang, H., Nagaswamy, U., Xia, Y., Gao, X., Fox, G.(2012) Biopolymers 97: 617
- PubMed: 22605553 Search on PubMed
- DOI: https://doi.org/10.1002/bip.22049
- Primary Citation Related Structures:
4A4R, 4A4S, 4A4T, 4A4U - PubMed Abstract:
The structures of four small RNAs each containing a different version of the UNAC loop were determined in solution using NMR spectroscopy and restrained molecular dynamics. The UMAC tetraloops (where M is A or C) exhibited a typical GNRA fold including at least one hydrogen bond between the first U and fourth C. In contrast, UGAC and UUAC tetraloops have a different orientation of the first and fourth residues, such that they do not closely mimic the GNRA fold. Although the UMAC tetraloops are excellent structural mimics of the GNRA tetraloop backbone, sequence comparisons typically do not reveal co-variation between the two loop types. The limited covariation is attributed to differences in the location of potential hydrogen bond donors and acceptors as a result of the replacement of the terminal A of GNRA with C in the UMAC version. Thus, UMAC loops do not readily form the common GNRA tetraloop-receptor interaction. The loop at positions 863-866 in E. coli 16S ribosomal RNA appears to be a major exception. However, in this case the GNRA loop does not in fact engage in the usual base to backbone tertiary interactions. In summary, UMAC loops are not just an alternative sequence version of the GNRA loop family, but instead they expand the types of interactions, or lack thereof, that are possible. From a synthetic biology perspective their inclusion in an artificial RNA may allow the establishment of a stable loop structure while minimizing unwanted long range interactions or permitting alternative long-range interactions. © 2012 Wiley Periodicals, Inc. Biopolymers 97: 617-628, 2012.
- Department of Biology and Biochemistry, University of Houston, Houston, TX, USA.
Organizational Affiliation: