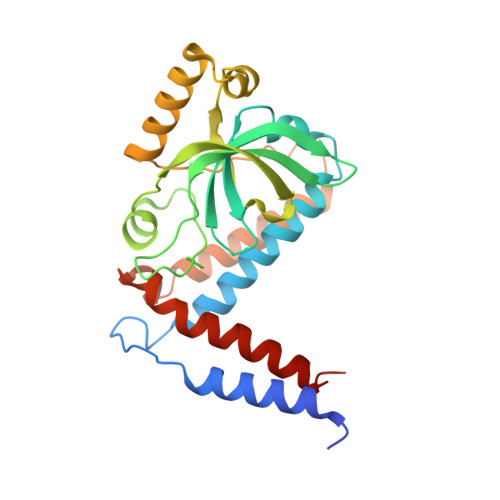
The Crystal Structure of S. Cerevisiae Ski2, a Dexh Helicase Associated with the Cytoplasmic Functions of the Exosome.
Halbach, F., Rode, M., Conti, E.(2012) RNA 18: 124
- PubMed: 22114319
- DOI: https://doi.org/10.1261/rna.029553.111
- Primary Citation Related Structures:
4A4K, 4A4Z - PubMed Abstract:
Ski2 is a cytoplasmic RNA helicase that functions together with the exosome in the turnover and quality control of mRNAs. Ski2 is conserved in eukaryotes and is related to the helicase Mtr4, a cofactor of the nuclear exosome involved in the processing and quality control of a variety of structured RNAs. We have determined the 2.4 Å resolution crystal structure of the 113 kDa helicase region of Saccharomyces cerevisiae Ski2. The structure shows that Ski2 has an overall architecture similar to that of Mtr4, with a core DExH region and an extended insertion domain. The insertion is not required for the formation of the Ski2-Ski3-Ski8 complex, but is instead an RNA-binding domain. While this is reminiscent of the Mtr4 insertion, there are specific structural and biochemical differences between the two helicases. The insertion of yeast Mtr4 consists of a β-barrel domain that is flexibly attached to a helical stalk, contains a KOW signature motif, and binds in vitro-transcribed tRNA(i)(Met), but not single-stranded RNA. The β-barrel domain of yeast Ski2 does not contain a KOW motif and is tightly packed against the helical stalk, forming a single structural unit maintained by a zinc-binding site. Biochemically, the Ski2 insertion has broad substrate specificity, binding both single-stranded and double-stranded RNAs. We speculate that the Ski2 and Mtr4 insertion domains have evolved with different properties tailored to the type of transcripts that are the substrates of the cytoplasmic and nuclear exosome.
- Department of Structural Cell Biology, Max-Planck-Institute of Biochemistry, D-82152 Martinsried, Germany.
Organizational Affiliation: