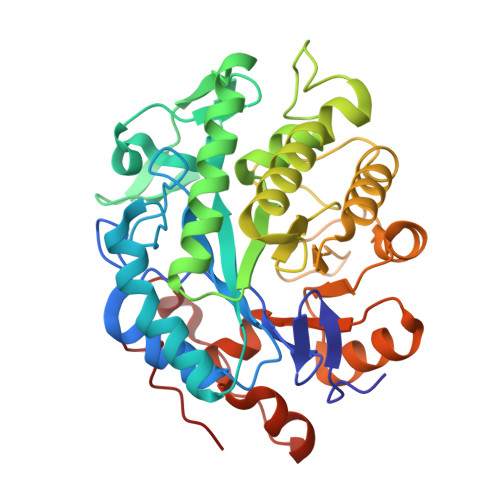
Crystal Structure Determination and Mutagenesis Analysis of the Ene Reductase Ncr.
Reich, S., Hoeffken, H.W., Rosche, B., Nestl, B.M., Hauer, B.(2012) Chembiochem 13: 2400
- PubMed: 23033175 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201200404
- Primary Citation Related Structures:
4A3U - PubMed Abstract:
The crystal structure of the "ene" nicotinamide-dependent cyclohexenone reductase (NCR) from Zymomonas mobilis (PDB ID: 4A3U) has been determined in complex with acetate ion, FMN, and nicotinamide, to a resolution of 1.95 Å. To study the activity and enantioselectivity of this enzyme in the bioreduction of activated α,β-unsaturated alkenes, the rational design methods site- and loop-directed mutagenesis were applied. Based on a multiple sequence alignment of various members of the Old Yellow Enzyme family, eight single-residue variants were generated and investigated in asymmetric bioreduction. Furthermore, a structural alignment of various ene reductases predicted four surface loop regions that are located near the entrance of the active site. Four NCR loop variants, derived from loop-swapping experiments with OYE1 from Saccharomyces pastorianus, were analysed for bioreduction. The three enzyme variants, P245Q, D337Y and F314Y, displayed increased activity compared to wild-type NCR towards the set of substrates tested. The active-site mutation Y177A demonstrated a clear influence on the enantioselectivity. The loop-swapping variants retained reduction efficiency, but demonstrated decreased enzyme activity compared with the wild-type NCR ene reductase enzyme.
- Universitaet Stuttgart, Institute of Technical Biochemistry, Allmandring 31, 70569 Stuttgart, Germany.
Organizational Affiliation: