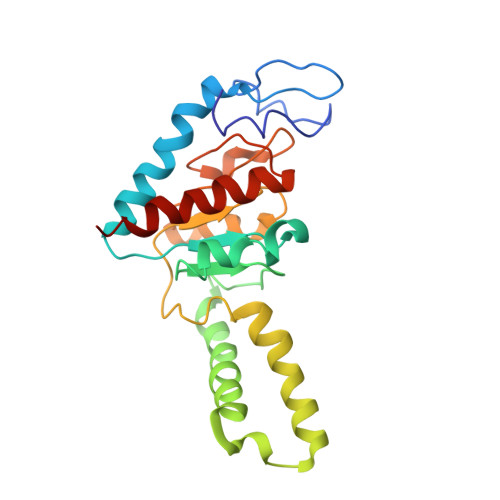
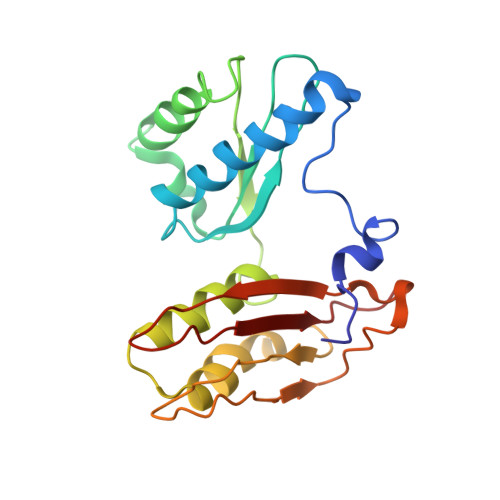
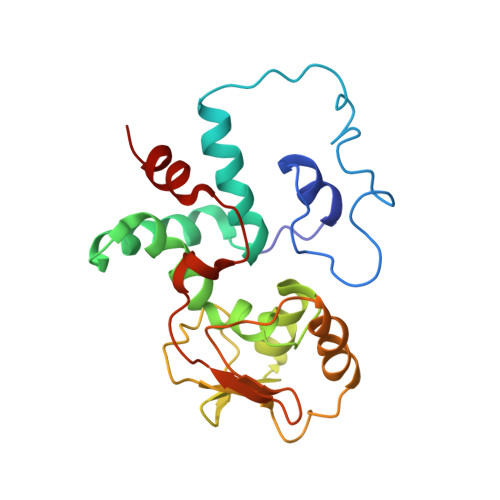
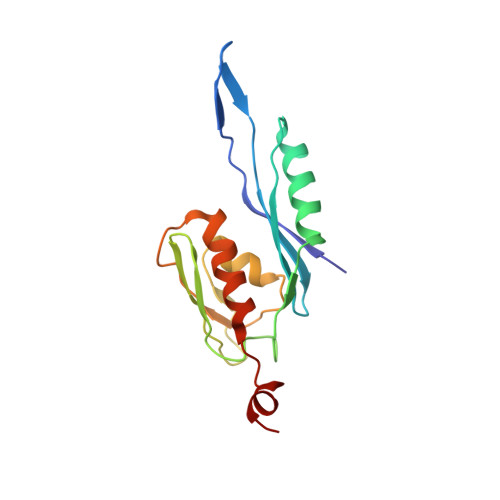
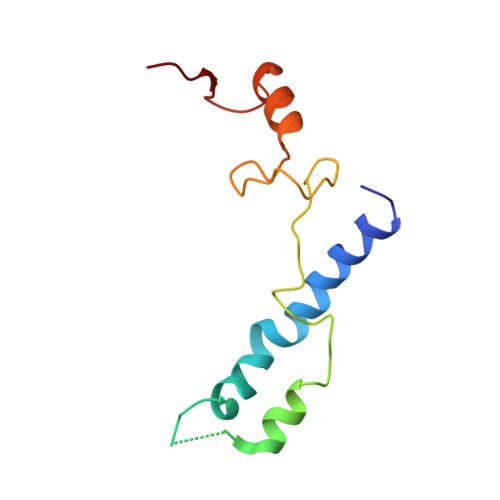
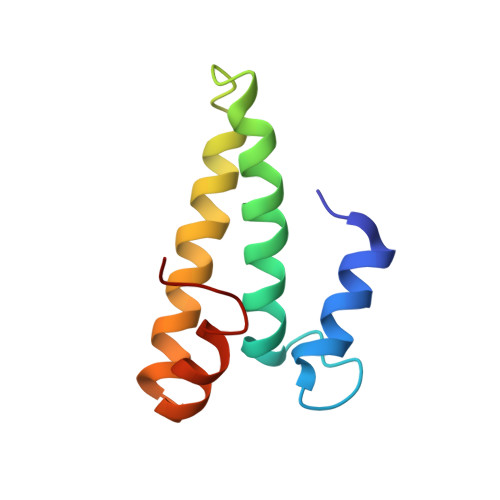
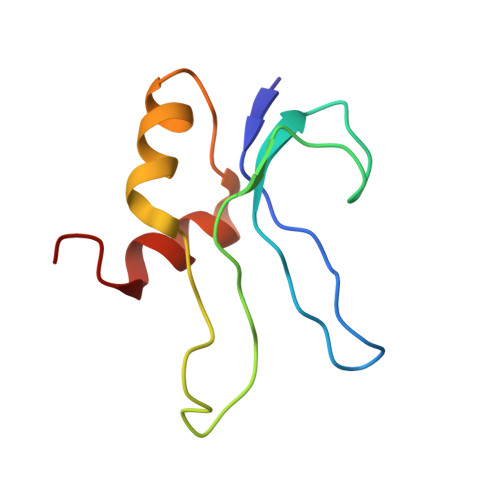
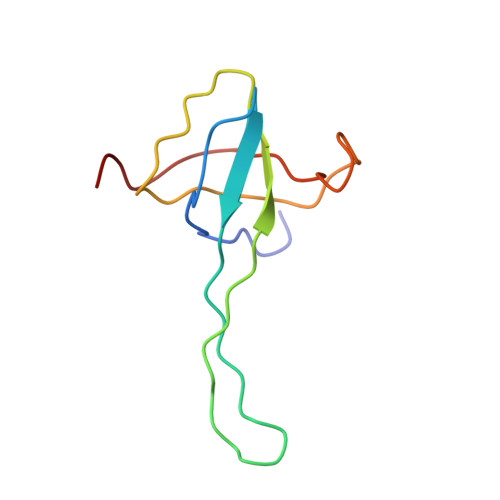
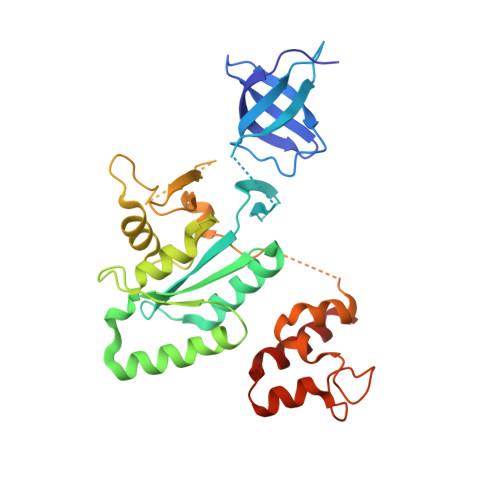
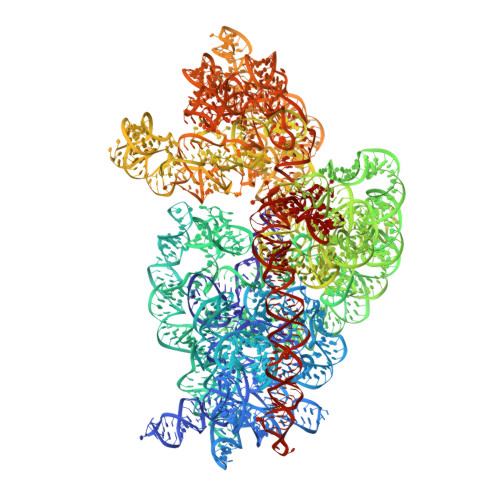
Cryo-Electron Microscopy Structure of the 30S Subunit in Complex with the Yjeq Biogenesis Factor.
Jomaa, A., Stewart, G., Mears, J.A., Kireeva, I., Brown, E.D., Ortega, J.(2011) RNA 17: 2026
- PubMed: 21960487 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.2922311
- Primary Citation Related Structures:
4A2I - PubMed Abstract:
YjeQ is a protein broadly conserved in bacteria containing an N-terminal oligonucleotide/oligosaccharide fold (OB-fold) domain, a central GTPase domain, and a C-terminal zinc-finger domain. YjeQ binds tightly and stoichiometrically to the 30S subunit, which stimulates its GTPase activity by 160-fold. Despite growing evidence for the involvement of the YjeQ protein in bacterial 30S subunit assembly, the specific function and mechanism of this protein remain unclear. Here, we report the costructure of YjeQ with the 30S subunit obtained by cryo-electron microscopy. The costructure revealed that YjeQ interacts simultaneously with helix 44, the head and the platform of the 30S subunit. This binding location of YjeQ in the 30S subunit suggests a chaperone role in processing of the 3' end of the rRNA as well as in mediating the correct orientation of the main domains of the 30S subunit. In addition, the YjeQ binding site partially overlaps with the interaction site of initiation factors 2 and 3, and upon binding, YjeQ covers three inter-subunit bridges that are important for the association of the 30S and 50S subunits. Hence, our structure suggests that YjeQ may assist in ribosome maturation by preventing premature formation of the translation initiation complex and association with the 50S subunit. Together, these results support a role for YjeQ in the late stages of 30S maturation.
- Department of Biochemistry and Biomedical Sciences and Michael G. DeGroote Institute for Infectious Diseases Research, McMaster University, Hamilton, Ontario, L8N3Z5, Canada.
Organizational Affiliation: