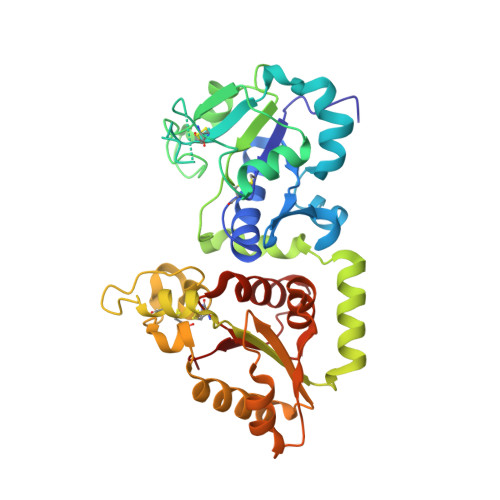
Structural Insights Into the Mechanism of Protein O-Fucosylation.
Lira-Navarrete, E., Valero-Gonzalez, J., Villanueva, R., Martinez-Julvez, M., Tejero, T., Merino, P., Panjikar, S., Hurtado-Guerrero, R.(2011) PLoS One 6: 25365
- PubMed: 21966509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0025365
- Primary Citation Related Structures:
3ZY2, 3ZY3, 3ZY4, 3ZY5, 3ZY6 - PubMed Abstract:
Protein O-fucosylation is an essential post-translational modification, involved in the folding of target proteins and in the role of these target proteins during embryonic development and adult tissue homeostasis, among other things. Two different enzymes are responsible for this modification, Protein O-fucosyltransferase 1 and 2 (POFUT1 and POFUT2, respectively). Both proteins have been characterised biologically and enzymatically but nothing is known at the molecular or structural level. Here we describe the first crystal structure of a catalytically functional POFUT1 in an apo-form and in complex with GDP-fucose and GDP. The enzyme belongs to the GT-B family and is not dependent on manganese for activity. GDP-fucose/GDP is localised in a conserved cavity connected to a large solvent exposed pocket, which we show is the binding site of epidermal growth factor (EGF) repeats in the extracellular domain of the Notch Receptor. Through both mutational and kinetic studies we have identified which residues are involved in binding and catalysis and have determined that the Arg240 residue is a key catalytic residue. We also propose a novel S(N)1-like catalytic mechanism with formation of an intimate ion pair, in which the glycosidic bond is cleaved before the nucleophilic attack; and theoretical calculations at a DFT (B3LYP/6-31+G(d,p) support this mechanism. Thus, the crystal structure together with our mutagenesis studies explain the molecular mechanism of POFUT1 and provide a new starting point for the design of functional inhibitors to this critical enzyme in the future.
- Institute of Biocomputation and Physics of Complex Systems, University of Zaragoza, Zaragoza, Spain.
Organizational Affiliation: