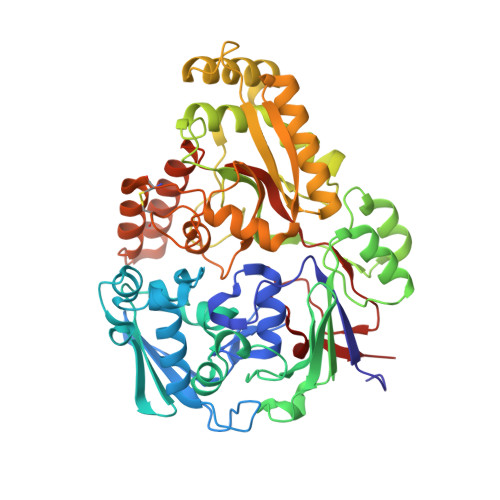
A Structure-Based Strategy for Epitope Discovery in Burkholderia Pseudomallei Oppa Antigen.
Lassaux, P., Peri, C., Ferrer-Navarro, M., Gourlay, L.J., Gori, A., Conchillo-Sole, O., Rinchai, D., Lertmemongkolchai, G., Longhi, R., Daura, X., Colombo, G., Bolognesi, M.(2013) Structure 21: 167
- PubMed: 23159127
- DOI: https://doi.org/10.1016/j.str.2012.10.005
- Primary Citation of Related Structures:
3ZS6 - PubMed Abstract:
We present an approach integrating structural and computational biology with immunological tests to identify epitopes in the OppA antigen from the Gram-negative pathogen Burkholderia pseudomallei, the etiological agent of melioidosis. The crystal structure of OppA(Bp), reported here at 2.1 Å resolution, was the basis for a computational analysis that identified three potential epitopes. In parallel, antigen proteolysis and immunocapturing allowed us to identify three additional peptides. All six potential epitopes were synthesized as free peptides and tested for their immunoreactivity against sera from healthy seronegative, healthy seropositive, and recovered melioidosis patients. Three synthetic peptides allowed the different patient groups to be distinguished, underlining the potential of this approach. Extension of the computational analysis, including energy-based decomposition methods, allowed rationalizing results of the predictive analyses and the immunocapture epitope mapping. Our results illustrate a structure-based epitope discovery process, whose application may expand our perspectives in the diagnostic and vaccine design fields.
- Department of Biosciences, University of Milan, Milan 20133, Italy.
Organizational Affiliation: