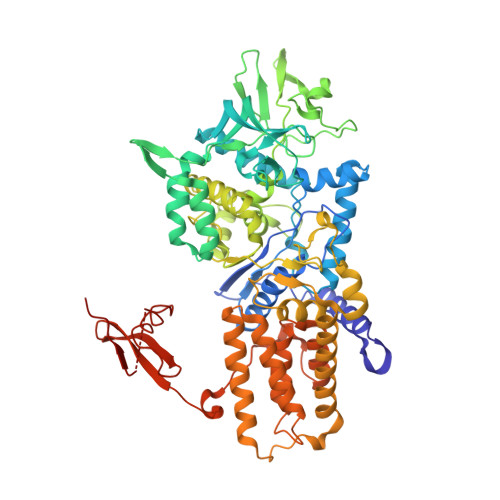
Leucyl-tRNA Synthetase Editing Domain Functions as a Molecular Rheostat to Control Codon Ambiguity in Mycoplasma Pathogens.
Li, L., Palencia, A., Lukk, T., Li, Z., Luthey-Schulten, Z.A., Cusack, S., Martinis, S.A., Boniecki, M.T.(2013) Proc Natl Acad Sci U S A 110: 3817
- PubMed: 23431144
- DOI: https://doi.org/10.1073/pnas.1218374110
- Primary Citation of Related Structures:
3ZIU - PubMed Abstract:
Mycoplasma leucyl-tRNA synthetases (LeuRSs) have been identified in which the connective polypeptide 1 (CP1) amino acid editing domain that clears mischarged tRNAs are missing (Mycoplasma mobile) or highly degenerate (Mycoplasma synoviae). Thus, these enzymes rely on a clearance pathway called pretransfer editing, which hydrolyzes misactivated aminoacyl-adenylate intermediate via a nebulous mechanism that has been controversial for decades. Even as the sole fidelity pathway for clearing amino acid selection errors in the pathogenic M. mobile, pretransfer editing is not robust enough to completely block mischarging of tRNA(Leu), resulting in codon ambiguity and statistical proteins. A high-resolution X-ray crystal structure shows that M. mobile LeuRS structurally overlaps with other LeuRS cores. However, when CP1 domains from different aminoacyl-tRNA synthetases and origins were fused to this common LeuRS core, surprisingly, pretransfer editing was enhanced. It is hypothesized that the CP1 domain evolved as a molecular rheostat to balance multiple functions. These include distal control of specificity and enzyme activity in the ancient canonical core, as well as providing a separate hydrolytic active site for clearing mischarged tRNA.
- Center for Biophysics and Computational Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA.
Organizational Affiliation: