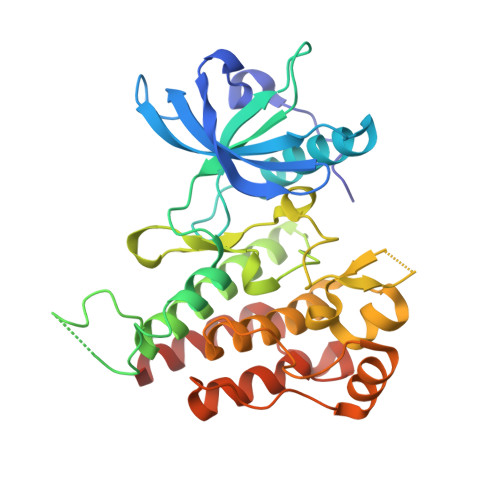
Distinct binding mode of multikinase inhibitor lenvatinib revealed by biochemical characterization.
Okamoto, K., Ikemori-Kawada, M., Jestel, A., von Konig, K., Funahashi, Y., Matsushima, T., Tsuruoka, A., Inoue, A., Matsui, J.(2015) ACS Med Chem Lett 6: 89-94
- PubMed: 25589937 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml500394m
- Primary Citation Related Structures:
3WZD, 3WZE - PubMed Abstract:
Lenvatinib is an oral multikinase inhibitor that selectively inhibits vascular endothelial growth factor (VEGF) receptors 1 to 3 and other proangiogenic and oncogenic pathway-related receptor tyrosine kinases. To elucidate the origin of the potency of lenvatinib in VEGF receptor 2 (VEGFR2) inhibition, we conducted a kinetic interaction analysis of lenvatinib with VEGFR2 and X-ray analysis of the crystal structure of VEGFR2-lenvatinib complexes. Kinetic analysis revealed that lenvatinib had a rapid association rate constant and a relatively slow dissociation rate constant in complex with VEGFR2. Co-crystal structure analysis demonstrated that lenvatinib binds at its ATP mimetic quinoline moiety to the ATP binding site and to the neighboring region via a cyclopropane ring, adopting an Asp-Phe-Gly (DFG)-"in" conformation. These results suggest that lenvatinib is very distinct in its binding mode of interaction compared to the several approved VEGFR2 kinase inhibitors.
- Eisai Co., Ltd , Tsukuba, Ibaraki 300-2635, Japan.
Organizational Affiliation: