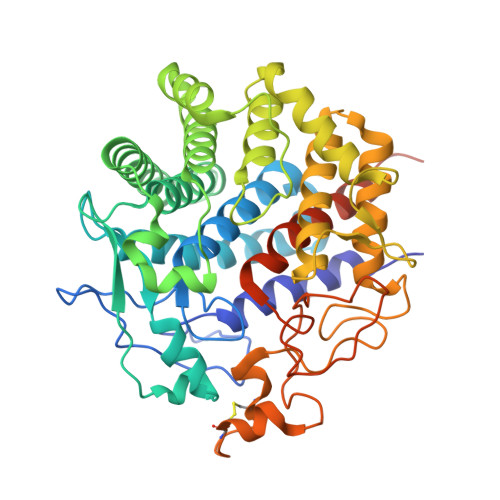
Crystal structure of endo-1,4-beta-glucanase from Eisenia fetida
Arimori, T., Ito, A., Nakazawa, M., Ueda, M., Tamada, T.(2013) J Synchrotron Radiat 20: 884-889
- PubMed: 24121333 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0909049513021110
- Primary Citation Related Structures:
3WC3 - PubMed Abstract:
The saccharification process is essential for bioethanol production from woody biomass including celluloses. Cold-adapted cellulase, which has sufficient activity at low temperature (<293 K), is capable of reducing heating costs during the saccharification process and is suitable for simultaneous saccharification and fermentation. Endo-1,4-β-glucanase from the earthworm Eisenia fetida (EF-EG2) belonging to glycoside hydrolase family 9 has been shown to have the highest activity at 313 K, and also retained a comparatively high activity at 283 K. The recombinant EF-EG2 was purified expressed in Pichia pastoris, and then grew needle-shaped crystals with dimensions of 0.02 × 0.02 × 1 mm. The crystals belonged to the space group P3221 with unit-cell parameters of a = b = 136 Å, c = 55.0 Å. The final model of EF-EG2, including 435 residues, two ions, seven crystallization reagents and 696 waters, was refined to a crystallographic R-factor of 14.7% (free R-factor of 16.8%) to 1.5 Å resolution. The overall structure of EF-EG2 has an (α/α)6 barrel fold which contains a putative active-site cleft and a negatively charged surface. This structural information helps us understand the catalytic and cold adaptation mechanisms of EF-EG2.
- Quantum Beam Science Directorate, Japan Atomic Energy Agency, 2-4 Shirakata-Shirane, Tokai, Ibaraki 319-1195, Japan.
Organizational Affiliation: