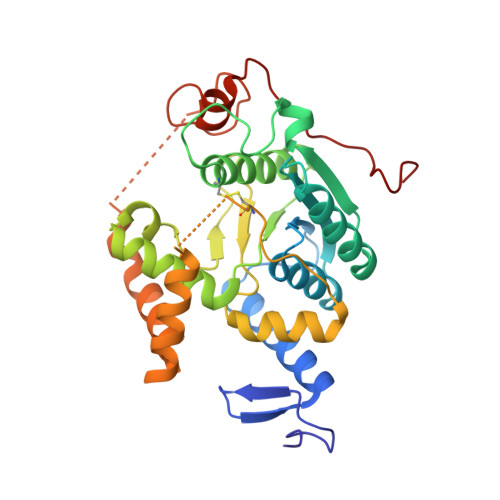
Crystallographic and mutational studies on the tRNA thiouridine synthetase TtuA.
Nakagawa, H., Kuratani, M., Goto-Ito, S., Ito, T., Katsura, K., Terada, T., Shirouzu, M., Sekine, S.I., Shigi, N., Yokoyama, S.(2013) Proteins
- PubMed: 23444054 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24273
- Primary Citation Related Structures:
3VRH - PubMed Abstract:
In thermophilic bacteria, specific 2-thiolation occurs on the conserved ribothymidine at position 54 (T54) in tRNAs, which is necessary for survival at high temperatures. T54 2-thiolation is achieved by the tRNA thiouridine synthetase TtuA and sulfur-carrier proteins. TtuA has five conserved CXXC/H motifs and the signature PP motif, and belongs to the TtcA family of tRNA 2-thiolation enzymes, for which there is currently no structural information. In this study, we determined the crystal structure of a TtuA homolog from the hyperthermophilic archeon Pyrococcus horikoshii at 2.1 Å resolution. The P. horikoshii TtuA forms a homodimer, and each subunit contains a catalytic domain and unique N- and C-terminal zinc fingers. The catalytic domain has much higher structural similarity to that of another tRNA modification enzyme, TilS (tRNA(Ile)₂ lysidine synthetase), than to the other type of tRNA 2-thiolation enzyme, MnmA. Three conserved cysteine residues are clustered in the putative catalytic site, which is not present in TilS. An in vivo mutational analysis in the bacterium Thermus thermophilus demonstrated that the three conserved cysteine residues and the putative ATP-binding residues in the catalytic domain are important for the TtuA activity. A positively charged surface that includes the catalytic site and the two zinc fingers is likely to provide the tRNA-binding site.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: