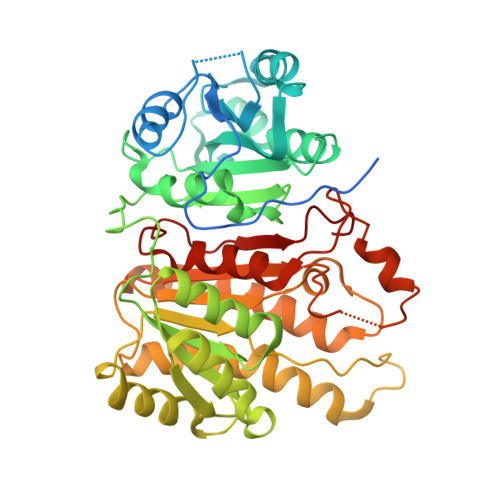
Crystal structure of hexanoyl-CoA bound to beta-ketoacyl reductase FabG4 of Mycobacterium tuberculosis
Dutta, D., Bhattacharyya, S., Roychowdhury, A., Biswas, R., Das, A.K.(2013) Biochem J 450: 127-139
- PubMed: 23163771
- DOI: https://doi.org/10.1042/BJ20121107
- Primary Citation of Related Structures:
3V1U, 4FW8 - PubMed Abstract:
FabGs, or β-oxoacyl reductases, are involved in fatty acid synthesis. The reaction entails NADPH/NADH-mediated conversion of β-oxoacyl-ACP (acyl-carrier protein) into β-hydroxyacyl-ACP. HMwFabGs (high-molecular-weight FabG) form a phylogenetically separate group of FabG enzymes. FabG4, an HMwFabG from Mycobacterium tuberculosis, contains two distinct domains, an N-terminal 'flavodoxintype' domain and a C-terminal oxoreductase domain. The catalytically active C-terminal domain utilizes NADH to reduce β-oxoacyl-CoA to β-hydroxyacyl-CoA. In the present study the crystal structures of the FabG4-NADH binary complex and the FabG4-NAD+-hexanoyl-CoA ternary complex have been determined to understand the substrate specificity and catalytic mechanism of FabG4. This is the first report to demonstrate how FabG4 interacts with its coenzyme NADH and hexanoyl-CoA that mimics an elongating fattyacyl chain covalently linked with CoA. Structural analysis shows that the binding of hexanoyl-CoA within the active site cavity of FabG significantly differs from that of the C16 fattyacyl substrate bound to mycobacterial FabI [InhA (enoyl-ACP reductase)]. The ternary complex reveals that both loop I and loop II interact with the phosphopantetheine moiety of CoA or ACP to align the covalently linked fattyacyl substrate near the active site. Structural data ACP inhibition studies indicate that FabG4 can accept both CoA- and ACP-based fattyacyl substrates. We have also shown that in the FabG4 dimer Arg146 and Arg445 of one monomer interact with the C-terminus of the second monomer to play pivotal role in substrate association and catalysis.
- Department of Biotechnology, Indian Institute of Technology Kharagpur, Kharagpur 721302, India.
Organizational Affiliation: