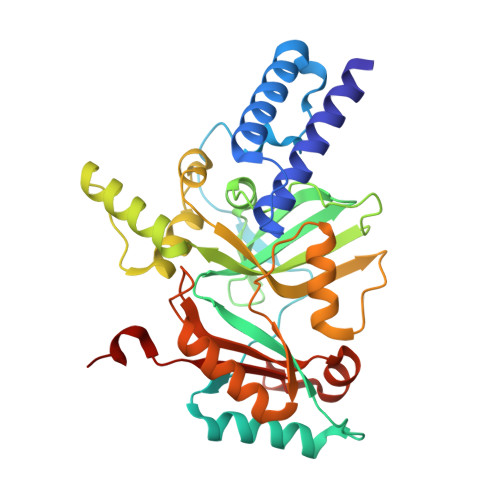
Structure Guided Understanding of NAD(+) Recognition in Bacterial DNA Ligases.
Lahiri, S.D., Gu, R.F., Gao, N., Karantzeni, I., Walkup, G.K., Mills, S.D.(2012) ACS Chem Biol 7: 571-580
- PubMed: 22230472
- DOI: https://doi.org/10.1021/cb200392g
- Primary Citation of Related Structures:
3UQ8 - PubMed Abstract:
NAD(+)-dependent DNA ligases (LigA) are essential bacterial enzymes that catalyze phosphodiester bond formation during DNA replication and repair processes. Phosphodiester bond formation proceeds through a 3-step reaction mechanism. In the first step, the LigA adenylation domain interacts with NAD(+) to form a covalent enzyme-AMP complex. Although it is well established that the specificity for binding of NAD(+) resides within the adenylation domain, the precise recognition elements for the initial binding event remain unclear. We report here the structure of the adenylation domain from Haemophilus influenzae LigA. This structure is a first snapshot of a LigA-AMP intermediate with NAD(+) bound to domain 1a in its open conformation. The binding affinities of NAD(+) for adenylated and nonadenylated forms of the H. influenzae LigA adenylation domain were similar. The combined crystallographic and NAD(+)-binding data suggest that the initial recognition of NAD(+) is via the NMN binding region in domain 1a of LigA.
- Department of Bioscience, Infection Innovative Medicines Unit, AstraZeneca R&D Boston, Waltham, Massachusetts 02451, United States. sushmita.lahiri@astrazeneca.com
Organizational Affiliation: