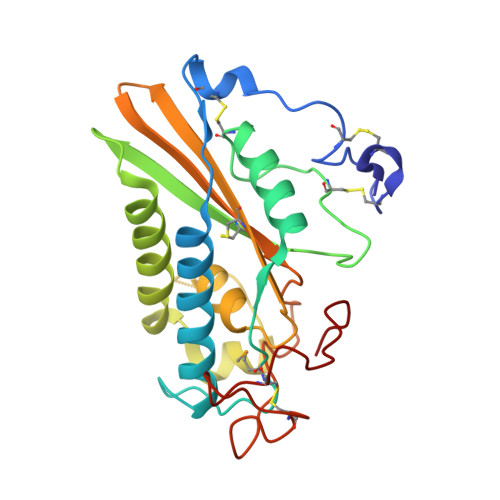
Structure of protein having inhibitory disintegrin and leukotriene scavenging functions contained in single domain.
Xu, X., Francischetti, I.M., Lai, R., Ribeiro, J.M., Andersen, J.F.(2012) J Biological Chem 287: 10967-10976
- PubMed: 22311975
- DOI: https://doi.org/10.1074/jbc.M112.340471
- Primary Citation of Related Structures:
3U3L, 3U3N, 3U3U - PubMed Abstract:
The antihemostatic/antiangiogenic protein tablysin-15 is a member of the CAP (cysteine-rich secretory, antigen 5, and pathogenesis-related 1 protein) superfamily and has been shown to bind the integrins α(IIb)β(3) and α(V)β(3) by means of an Arg-Gly-Asp (RGD) tripeptide sequence. Here we describe the x-ray crystal structure of tablysin-15 and show that the RGD motif is located in a novel structural context. The motif itself is contained in a type II β-turn structure that is similar in its conformation to the RGD sequence of the cyclic pentapeptide cilengitide when bound to integrin α(V)β(3). The CAP domain also contains a hydrophobic channel that appears to bind a fatty acid molecule in the crystal structure after purification from Escherichia coli. After delipidation of the protein, tablysin-15 was found to bind proinflammatory cysteinyl leukotrienes with submicromolar affinities. The structure of the leukotriene E(4)-tablysin-15 complex shows that the ligand binds with the nonfunctionalized end of the fatty acid chain buried in the hydrophobic pocket, whereas the carboxylate end of the ligand binds forms hydrogen bond/salt bridge interactions with polar side chains at the channel entrance. Therefore, tablysin-15 functions as an inhibitor of integrin function and as an anti-inflammatory scavenger of eicosanoids.
- Laboratory of Malaria and Vector Research, NIAID, National Intitutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: