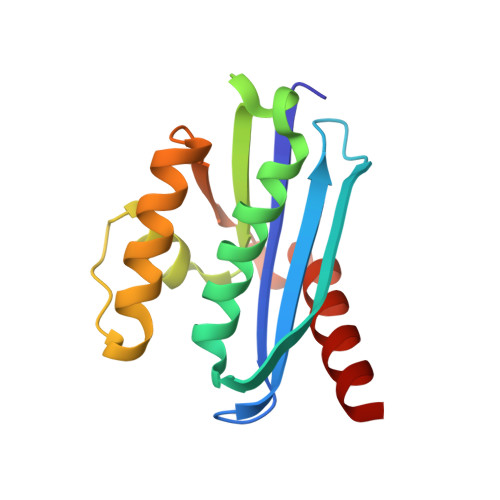
Activity, stability, and structure of metagenome-derived LC11-RNase H1, a homolog of Sulfolobus tokodaii RNase H1
Nguyen, T.N., Angkawidjaja, C., Kanaya, E., Koga, Y., Takano, K., Kanaya, S.(2012) Protein Sci 21: 553-561
- PubMed: 22389131
- DOI: https://doi.org/10.1002/pro.2043
- Primary Citation of Related Structures:
3U3G - PubMed Abstract:
Metagenome-derived LC11-RNase H1 is a homolog of Sulfolobus tokodaii RNase H1 (Sto-RNase H1). It lacks a C-terminal tail, which is responsible for hyperstabilization of Sto-RNase H1. Sto-RNase H1 is characterized by its ability to cleave not only an RNA/DNA hybrid but also a double-stranded RNA (dsRNA). To examine whether LC11-RNase H1 also exhibits both RNase H and dsRNase activities, LC11-RNase H1 was overproduced in Escherichia coli, purified, and characterized. LC11-RNase H1 exhibited RNase H activity with similar metal ion preference, optimum pH, and cleavage mode of substrate with those of Sto-RNase H1. However, LC11-RNase H1 did not exhibit dsRNase activity at any condition examined. LC11-RNase H1 was less stable than Sto-RNases H1 and its derivative lacking the C-terminal tail (Sto-RNase H1ΔC6) by 37 and 13 °C in T(m) , respectively. To understand the structural bases for these differences, the crystal structure of LC11-RNase H1 was determined at 1.4 Å resolution. The LC11-RNase H1 structure is highly similar to the Sto-RNase H1 structure. However, LC11-RNase H1 has two grooves on protein surface, one containing the active site and the other containing DNA-phosphate binding pocket, while Sto-RNase H1 has one groove containing the active site. In addition, LC11-RNase H1 contains more cavities and buried charged residues than Sto-RNase H1. We propose that LC11-RNase H1 does not exhibit dsRNase activity because dsRNA cannot fit to the two grooves on protein surface and that LC11-RNase H1 is less stable than Sto-RNase H1ΔC6 because of the increase in cavity volume and number of buried charged residues.
- Department of Material and Life Science, Graduate School of Engineering, Osaka University, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: