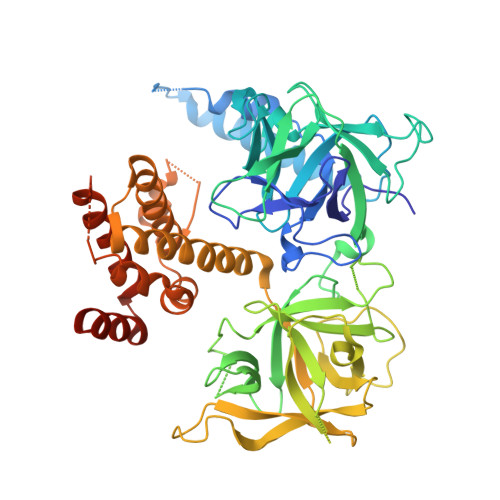
Apo and InsP(3)-bound crystal structures of the ligand-binding domain of an InsP(3) receptor.
Lin, C.C., Baek, K., Lu, Z.(2011) Nat Struct Mol Biol 18: 1172-1174
- PubMed: 21892169
- DOI: https://doi.org/10.1038/nsmb.2112
- Primary Citation of Related Structures:
3T8S - PubMed Abstract:
We report the crystal structures of the ligand-binding domain (LBD) of a rat inositol 1,4,5-trisphosphate receptor (InsP(3)R) in its apo and InsP(3)-bound conformations. Comparison of these two conformations reveals that LBD's first β-trefoil fold (β-TF1) and armadillo repeat fold (ARF) move together as a unit relative to its second β-trefoil fold (β-TF2). Whereas apo LBD may spontaneously transition between gating conformations, InsP(3) binding shifts this equilibrium toward the active state.
- Department of Physiology, Howard Hughes Medical Institute, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Organizational Affiliation: