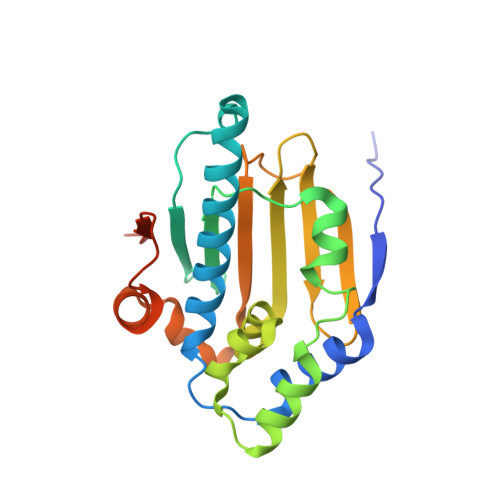
Structure insights into mechanisms of ATP hydrolysis and the activation of human heat-shock protein 90.
Li, J., Sun, L., Xu, C., Yu, F., Zhou, H., Zhao, Y., Zhang, J., Cai, J., Mao, C., Tang, L., Xu, Y., He, J.(2012) Acta Biochim Biophys Sin (Shanghai) 44: 300-306
- PubMed: 22318716 Search on PubMed
- DOI: https://doi.org/10.1093/abbs/gms001
- Primary Citation Related Structures:
3T0H, 3T0Z, 3T10 - PubMed Abstract:
The activation of molecular chaperone heat-shock protein 90 (Hsp90) is dependent on ATP binding and hydrolysis, which occurs in the N-terminal domains of protein. Here, we have determined three crystal structures of the N-terminal domain of human Hsp90 in native and in complex with ATP and ATP analog, providing a clear view of the catalytic mechanism of ATP hydrolysis by Hsp90. Additionally, the binding of ATP leads the N-terminal domains to be an intermediate state that could be used to partially explain why the isolated N-terminal domain of Hsp90 has very weak ATP hydrolytic activity.
- Shanghai Institute of Applied Physics, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: