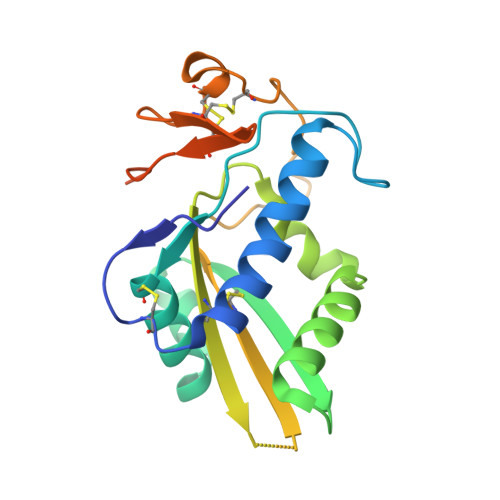
Hookworm SCP/TAPS protein structure--A key to understanding host-parasite interactions and developing new interventions.
Osman, A., Wang, C.K., Winter, A., Loukas, A., Tribolet, L., Gasser, R.B., Hofmann, A.(null) Biotechnol Adv 30: 652-657
- PubMed: 22120067
- DOI: https://doi.org/10.1016/j.biotechadv.2011.11.002
- Primary Citation Related Structures:
3S6S, 3S6U, 3S6V - PubMed Abstract:
SCP/TAPS proteins are a diverse family of molecules in eukaryotes, including parasites. Despite their abundant occurrence in parasite secretomes, very little is known about their functions in parasitic nematodes, including blood-feeding hookworms. Current information indicates that SCP/TAPS proteins (called Ancylostoma-secreted proteins, ASPs) of the canine hookworm, Ancylostoma caninum, represent at least three distinct groups of proteins. This information, combined with comparative modelling, indicates that all known ASPs have an equatorial groove that binds extended structures, such as peptides or glycans. To elucidate structure-function relationships, we explored the three-dimensional crystal structure of an ASP (called Ac-ASP-7), which is highly up-regulated in expression in the transition of A. caninum larvae from a free-living to a parasitic stage. The topology of the N-terminal domain is consistent with pathogenesis-related proteins, and the C-terminal extension that resembles the fold of the Hinge domain. By anomalous diffraction, we identified a new metal binding site in the C-terminal extension of the protein. Ac-ASP-7 is in a monomer-dimer equilibrium, and crystal-packing analysis identified a dimeric structure which might resemble the homo-dimer in solution. The dimer interaction interface includes a novel binding site for divalent metal ions, and is proposed to serve as a binding site for proteins involved in the parasite-host interplay at the molecular level. Understanding this interplay and the integration of structural and functional data could lead to the design of new approaches for the control of parasitic diseases, with biotechnological outcomes.
- Structural Chemistry Program, Eskitis Institute for Cell & Molecular Therapies, Griffith University, Brisbane, Queensland, Australia.
Organizational Affiliation: