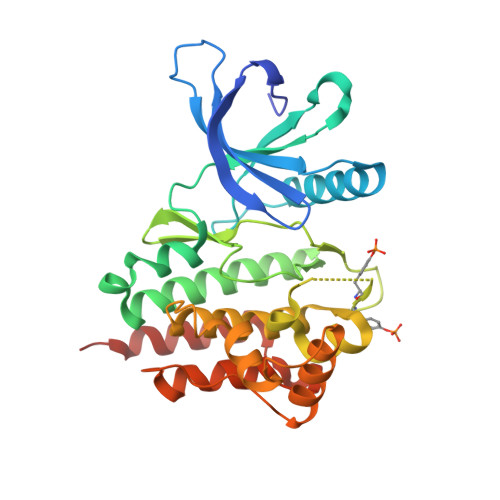
Discovery of 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitors of Janus kinase 2 (JAK2) for the treatment of myeloproliferative disorders.
Lim, J., Taoka, B., Otte, R.D., Spencer, K., Dinsmore, C.J., Altman, M.D., Chan, G., Rosenstein, C., Sharma, S., Su, H.P., Szewczak, A.A., Xu, L., Yin, H., Zugay-Murphy, J., Marshall, C.G., Young, J.R.(2011) J Med Chem 54: 7334-7349
- PubMed: 21942426 Search on PubMed
- DOI: https://doi.org/10.1021/jm200909u
- Primary Citation Related Structures:
3RVG - PubMed Abstract:
The JAK-STAT pathway mediates signaling by cytokines, which control survival, proliferation, and differentiation of a variety of cells. In recent years, a single point mutation (V617F) in the tyrosine kinase JAK2 was found to be present with a high incidence in myeloproliferative disorders (MPDs). This mutation led to hyperactivation of JAK2, cytokine-independent signaling, and subsequent activation of downstream signaling networks. The genetic, biological, and physiological evidence suggests that JAK2 inhibitors could be effective in treating MPDs. De novo design efforts of new scaffolds identified 1-amino-5H-pyrido[4,3-b]indol-4-carboxamides as a new viable lead series. Subsequent optimization of cell potency, metabolic stability, and off-target activities of the leads led to the discovery of 7-(2-aminopyrimidin-5-yl)-1-{[(1R)-1-cyclopropyl-2,2,2-trifluoroethyl]amino}-5H-pyrido[4,3-b]indole-4-carboxamide (65). Compound 65 is a potent, orally active inhibitor of JAK2 with excellent selectivity, PK profile, and in vivo efficacy in animal models.
- Department of Chemistry, Merck & Co., Inc., 33 Avenue Louis Pasteur, Boston, Massachusetts 02115, United States. jongwon_lim@merck.com
Organizational Affiliation: