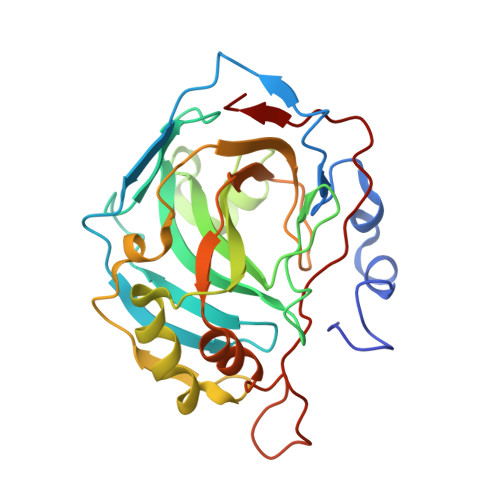
Structure and catalysis by carbonic anhydrase II: role of active-site tryptophan 5.
Mikulski, R., Domsic, J.F., Ling, G., Tu, C., Robbins, A.H., Silverman, D.N., McKenna, R.(2011) Arch Biochem Biophys 516: 97-102
- PubMed: 22001224 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.abb.2011.09.011
- Primary Citation Related Structures:
3RG3, 3RG4, 3RGE - PubMed Abstract:
The tryptophan residue Trp5, highly conserved in the α class of carbonic anhydrases including human carbonic anhydrase II (HCA II), is positioned at the entrance of the active site cavity and forms a π-stacking interaction with the imidazole ring of the proton shuttle His64 in its outward orientation. We have observed that replacement of Trp5 in HCA II caused significant structural changes, as determined by X-ray diffraction, in the conformation of 11 residues at the N-terminus and in the orientation of the proton shuttle residue His64. Most significantly, two variants W5H and W5E HCA II had His64 predominantly outward in orientation, while W5F and wild type showed the superposition of both outward and inward orientations in crystal structures. Although Trp5 influences the orientation of the proton shuttle His64, this orientation had no significant effect on the rate constant for proton transfer near 1μs(-1), determined by exchange of (18)O between CO(2) and water measured by mass spectrometry. The apparent values of the pK(a) of the zinc-bound water and the proton shuttle residue suggest that different active-site conformations influence the two stages of catalysis, the proton transfer stage and the interconversion of CO(2) and bicarbonate.
- Department of Pharmacology and Therapeutics, University of Florida, Gainesville, United States.
Organizational Affiliation: