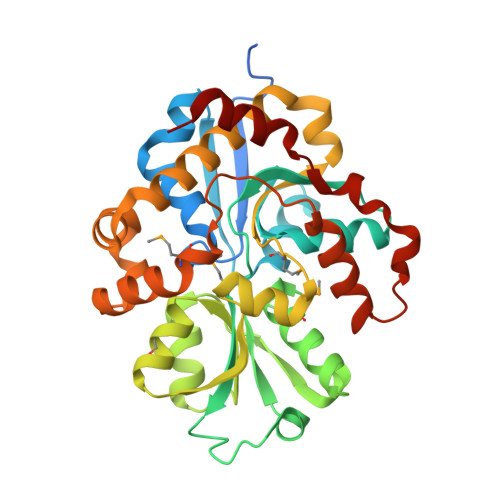
The crystal structure of pyrimidine/thiamin biosynthesis precursor-like domain-containing protein CAE31940 from proteobacterium Bordetella bronchiseptica RB50, and evolutionary insight into the NMT1/THI5 family.
Bajor, J., Tkaczuk, K.L., Chruszcz, M., Chapman, H., Kagan, O., Savchenko, A., Minor, W.(2014) J Struct Funct Genomics 15: 73-81
- PubMed: 24908050 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s10969-014-9180-3
- Primary Citation Related Structures:
3QSL - PubMed Abstract:
We report a 2.0 Å structure of the CAE31940 protein, a proteobacterial NMT1/THI5-like domain-containing protein. We also discuss the primary and tertiary structure similarity with its homologs. The highly conserved FGGXMP motif was identified in CAE31940, which corresponds to the GCCCX motif located in the vicinity of the active center characteristic for THi5-like proteins found in yeast. This suggests that the FGGXMP motif may be a unique hallmark of proteobacterial NMT1/THI5-like proteins.
- Department of Molecular Physiology and Biological Physics, University of Virginia, 1340 Jefferson Park Avenue, Charlottesville, VA, 22908, USA.
Organizational Affiliation: