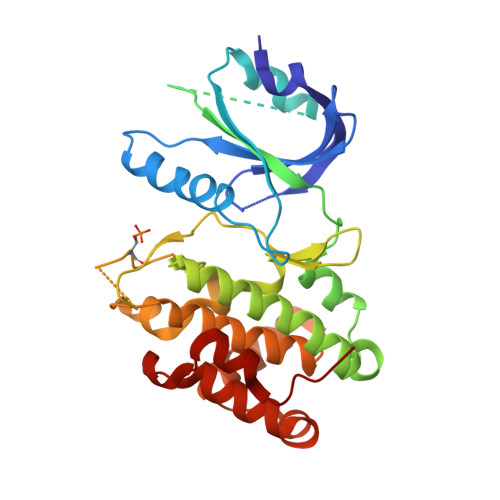
The structure of the PERK kinase domain suggests the mechanism for its activation.
Cui, W., Li, J., Ron, D., Sha, B.(2011) Acta Crystallogr D Biol Crystallogr 67: 423-428
- PubMed: 21543844 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444911006445
- Primary Citation Related Structures:
3QD2 - PubMed Abstract:
The endoplasmic reticulum (ER) unfolded protein response (UPR) is comprised of several intracellular signaling pathways that alleviate ER stress. The ER-localized transmembrane kinase PERK is one of three major ER stress transducers. Oligomerization of PERK's N-terminal ER luminal domain by ER stress promotes PERK trans-autophosphorylation of the C-terminal cytoplasmic kinase domain at multiple residues including Thr980 on the kinase activation loop. Activated PERK phosphorylates Ser51 of the α-subunit of translation initiation factor 2 (eIF2α), which inhibits initiation of protein synthesis and reduces the load of unfolded proteins entering the ER. The crystal structure of PERK's kinase domain has been determined to 2.8 Å resolution. The structure resembles the back-to-back dimer observed in the related eIF2α kinase PKR. Phosphorylation of Thr980 stabilizes both the activation loop and helix αG in the C-terminal lobe, preparing the latter for eIF2α binding. The structure suggests conservation in the mode of activation of eIF2α kinases and is consistent with a `line-up' model for PERK activation triggered by oligomerization of its luminal domain.
- Department of Cell Biology, University of Alabama at Birmingham, Birmingham, AL 35294, USA.
Organizational Affiliation: