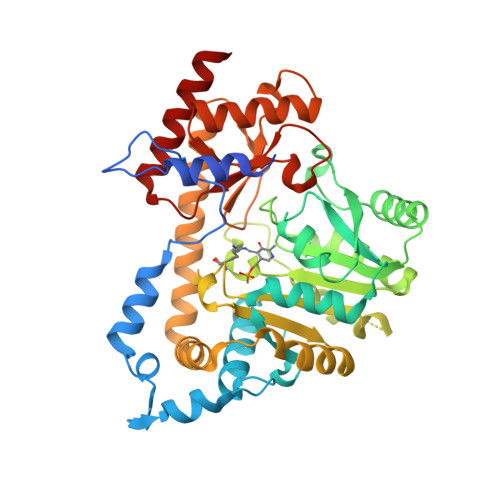
Structural basis for reduced activity of 1-aminocyclopropane-1-carboxylate synthase affected by a mutation linked to andromonoecy.
Scharer, M.A., Eliot, A.C., Grutter, M.G., Capitani, G.(2011) FEBS Lett 585: 111-114
- PubMed: 21075107 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2010.11.013
- Primary Citation Related Structures:
3PIU - PubMed Abstract:
1-aminocyclopropane-1-carboxylate synthase (ACS) is a key enzyme in the biosynthesis of the plant hormone ethylene. Recently, a new biological role for ACS has been found in Cucumis melo where a single point mutation (A57V) of one isoform of the enzyme, causing reduced activity, results in andromonoecious plants. We present here a straightforward structural basis for the reduced activity of the A57V mutant, based on our work on Malus domestica ACS, including a new structure of the unliganded apple enzyme at 1.35Å resolution.
- Biomolecular Research, Paul Scherrer Institut, Villigen PSI, Switzerland.
Organizational Affiliation: