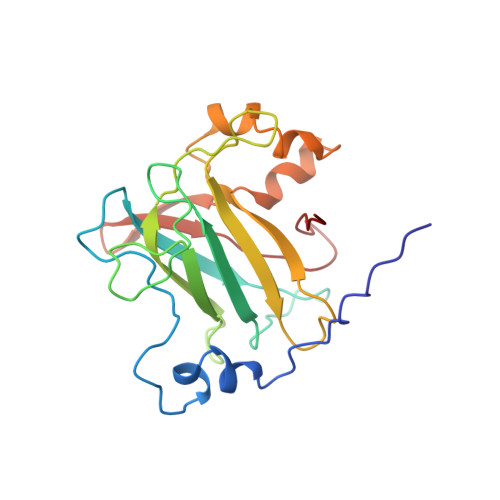
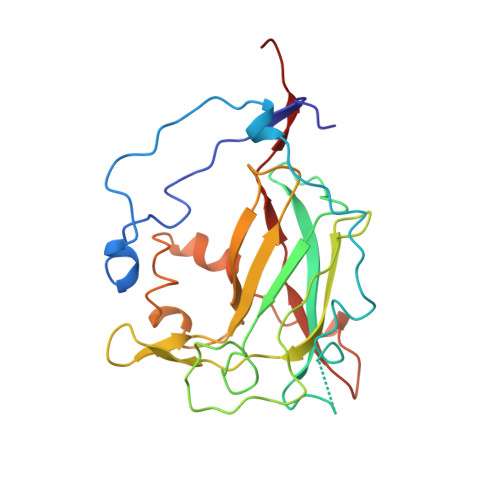
Structures of competitive inhibitor complexes of protocatechuate 3,4-dioxygenase: multiple exogenous ligand binding orientations within the active site.
Orville, A.M., Elango, N., Lipscomb, J.D., Ohlendorf, D.H.(1997) Biochemistry 36: 10039-10051
- PubMed: 9254599 Search on PubMed
- DOI: https://doi.org/10.1021/bi970468n
- Primary Citation Related Structures:
3PCB, 3PCC, 3PCE, 3PCF, 3PCG, 3PCH, 3PCI - PubMed Abstract:
Protocatechuate 3,4-dioxygenase (3,4-PCD) catalyzes the oxidative ring cleavage of 3,4-dihydroxybenzoate to produce beta-carboxy-cis, cis-muconate. Crystal structures of Pseudomonas putida3,4-PCD [quaternary structure of (alphabetaFe3+)12] complexed with seven competitive inhibitors [3-hydroxyphenylacetate (MHP), 4-hydroxyphenylacetate (PHP), 3-hydroxybenzoate (MHB), 4-hydroxybenzoate (PHB), 3-fluoro-4-hydroxybenzoate (FHB), 3-chloro-4-hydroxybenzoate (CHB), and 3-iodo-4-hydroxybenzoate (IHB)] are reported at 2.0-2.2 A resolution with R-factors of 0. 0.159-0.179. The inhibitors bind in a narrow active site crevasse lined with residues that provide a microenvironment that closely matches the chemical characteristics of the inhibitors. This results in as little as 20% solvent-exposed surface area for the higher-affinity inhibitors (PHB, CHB, and FHB). In uncomplexed 3,4-PCD, the active site Fe3+ is bound at the bottom of the active site crevasse by four endogenous ligands and a solvent molecule (Wat827). The orientations of the endogenous ligands are relatively unperturbed in each inhibitor complex, but the inhibitors themselves bind to or near the iron in a range of positions, all of which perturb the position of Wat827. The three lowest-affinity inhibitors (MHP, PHP, and IHB) yield distorted trigonal bipyramidal iron coordination geometry in which the inhibitor C4-phenolate group displaces the solvent ligand. MHB binds within the active site, but neither its C3-OH group nor the solvent molecule binds to the iron. The C4-phenolate group of the three highest-affinity inhibitors (PHB, CHB, and FHB) coordinates the Fe3+ adjacent to Wat827, resulting in a shift in its position to yield a six-coordinate distorted octahedral geometry. The range of inhibitor orientations may mimic the mechanistically significant stages of substrate binding to 3, 4-PCD. The structure of the final substrate complex is reported in the following paper [Orville, A. M., Lipscomb, J. D., & Ohlendorf, D. H. (1997) Biochemistry 36, 10052-10066].
- Department of Biochemistry, Medical School, and Center for Metals in Biocatalysis, University of Minnesota, Minneapolis, Minnesota 55455-0347, USA.
Organizational Affiliation: