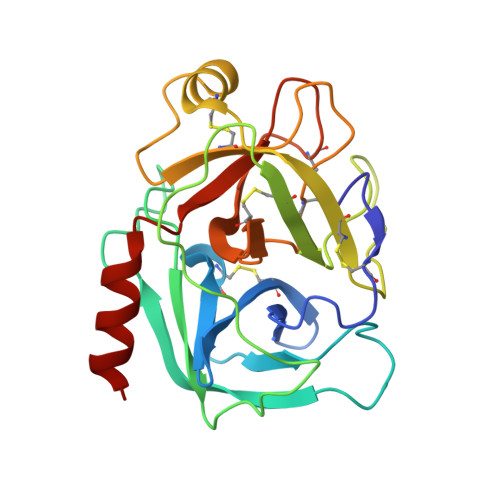
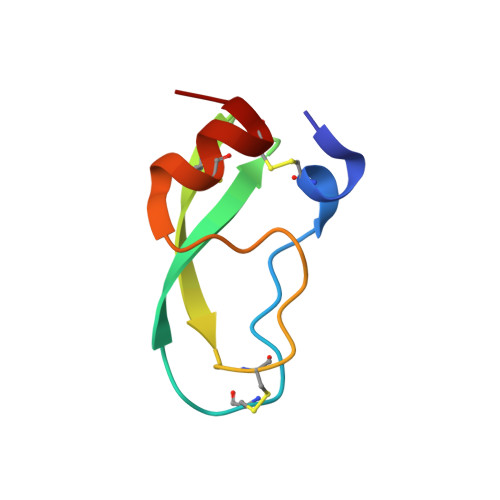
The P2' residue is a key determinant of mesotrypsin specificity: engineering a high-affinity inhibitor with anticancer activity.
Salameh, M.A., Soares, A.S., Hockla, A., Radisky, D.C., Radisky, E.S.(2011) Biochem J 440: 95-105
- PubMed: 21806544 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20110788
- Primary Citation Related Structures:
3P92, 3P95 - PubMed Abstract:
PRSS3/mesotrypsin is an atypical isoform of trypsin, the up-regulation of which has been implicated in promoting tumour progression. Mesotrypsin inhibitors could potentially provide valuable research tools and novel therapeutics, but small-molecule trypsin inhibitors have low affinity and little selectivity, whereas protein trypsin inhibitors bind poorly and are rapidly degraded by mesotrypsin. In the present study, we use mutagenesis of a mesotrypsin substrate, APPI (amyloid precursor protein Kunitz protease inhibitor domain), and of a poor mesotrypsin inhibitor, BPTI (bovine pancreatic trypsin inhibitor), to dissect mesotrypsin specificity at the key P(2)' position. We find that bulky and charged residues strongly disfavour binding, whereas acidic residues facilitate catalysis. Crystal structures of mesotrypsin complexes with BPTI variants provide structural insights into mesotrypsin specificity and inhibition. Through optimization of the P(1) and P(2)' residues of BPTI, we generate a stable high-affinity mesotrypsin inhibitor with an equilibrium binding constant K(i) of 5.9 nM, a >2000-fold improvement in affinity over native BPTI. Using this engineered inhibitor, we demonstrate the efficacy of pharmacological inhibition of mesotrypsin in assays of breast cancer cell malignant growth and pancreatic cancer cell invasion. Although further improvements in inhibitor selectivity will be important before clinical potential can be realized, the results of the present study support the feasibility of engineering protein protease inhibitors of mesotrypsin and highlight their therapeutic potential.
- Department of Cancer Biology, Mayo Clinic Cancer Center, Jacksonville, FL 32224, USA.
Organizational Affiliation: