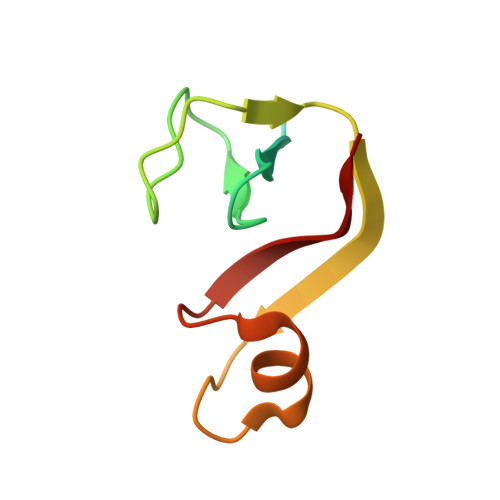
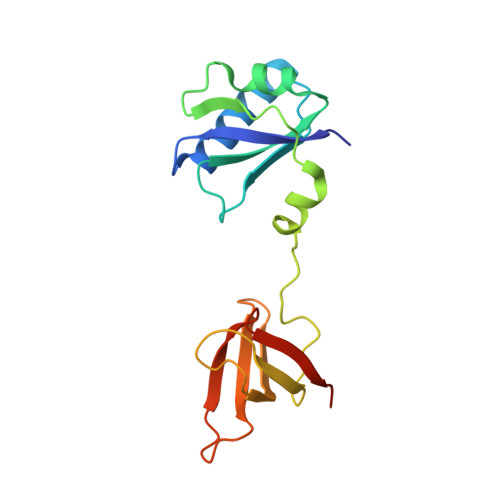
RNA polymerase and transcription elongation factor Spt4/5 complex structure.
Klein, B.J., Bose, D., Baker, K.J., Yusoff, Z.M., Zhang, X., Murakami, K.S.(2011) Proc Natl Acad Sci U S A 108: 546-550
- PubMed: 21187417 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1013828108
- Primary Citation Related Structures:
3P8B - PubMed Abstract:
Spt4/5 in archaea and eukaryote and its bacterial homolog NusG is the only elongation factor conserved in all three domains of life and plays many key roles in cotranscriptional regulation and in recruiting other factors to the elongating RNA polymerase. Here, we present the crystal structure of Spt4/5 as well as the structure of RNA polymerase-Spt4/5 complex using cryoelectron microscopy reconstruction and single particle analysis. The Spt4/5 binds in the middle of RNA polymerase claw and encloses the DNA, reminiscent of the DNA polymerase clamp and ring helicases. The transcription elongation complex model reveals that the Spt4/5 is an upstream DNA holder and contacts the nontemplate DNA in the transcription bubble. These structures reveal that the cellular RNA polymerases also use a strategy of encircling DNA to enhance its processivity as commonly observed for many nucleic acid processing enzymes including DNA polymerases and helicases.
- Department of Biochemistry and Molecular Biology, Pennsylvania State University, University Park, PA 16802, USA.
Organizational Affiliation: