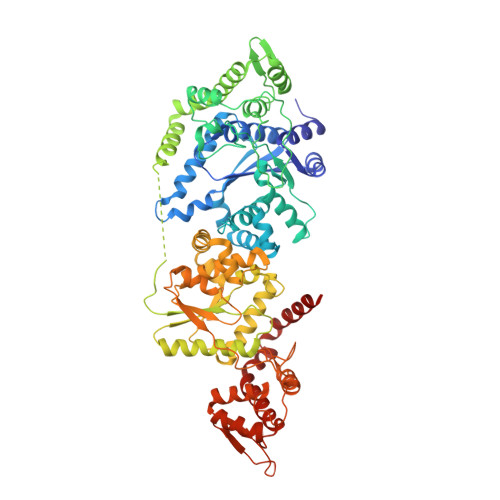
The Double-Length Tyrosyl-tRNA Synthetase from the Eukaryote Leishmania major Forms an Intrinsically Asymmetric Pseudo-Dimer.
Larson, E.T., Kim, J.E., Castaneda, L.J., Napuli, A.J., Zhang, Z., Fan, E., Zucker, F.H., Verlinde, C.L., Buckner, F.S., Van Voorhis, W.C., Hol, W.G., Merritt, E.A.(2011) J Mol Biology 409: 159-176
- PubMed: 21420975
- DOI: https://doi.org/10.1016/j.jmb.2011.03.026
- Primary Citation of Related Structures:
3P0H, 3P0I, 3P0J - PubMed Abstract:
The single tyrosyl-tRNA synthetase (TyrRS) gene in trypanosomatid genomes codes for a protein that is twice the length of TyrRS from virtually all other organisms. Each half of the double-length TyrRS contains a catalytic domain and an anticodon-binding domain; however, the two halves retain only 17% sequence identity to each other. The structural and functional consequences of this duplication and divergence are unclear. TyrRS normally forms a homodimer in which the active site of one monomer pairs with the anticodon-binding domain from the other. However, crystal structures of Leishmania major TyrRS show that, instead, the two halves of a single molecule form a pseudo-dimer resembling the canonical TyrRS dimer. Curiously, the C-terminal copy of the catalytic domain has lost the catalytically important HIGH and KMSKS motifs characteristic of class I aminoacyl-tRNA synthetases. Thus, the pseudo-dimer contains only one functional active site (contributed by the N-terminal half) and only one functional anticodon recognition site (contributed by the C-terminal half). Despite biochemical evidence for negative cooperativity between the two active sites of the usual TyrRS homodimer, previous structures have captured a crystallographically-imposed symmetric state. As the L. major TyrRS pseudo-dimer is inherently asymmetric, conformational variations observed near the active site may be relevant to understanding how the state of a single active site is communicated across the dimer interface. Furthermore, substantial differences between trypanosomal TyrRS and human homologs are promising for the design of inhibitors that selectively target the parasite enzyme.
- Medical Structural Genomics of Pathogenic Protozoa, Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: