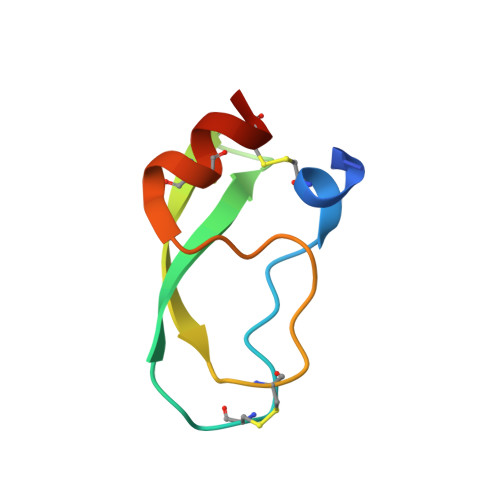
Structure of the recombinant BPTI/Kunitz-type inhibitor rShPI-1A from the marine invertebrate Stichodactyla helianthus.
Garcia-Fernandez, R., Pons, T., Meyer, A., Perbandt, M., Gonzalez-Gonzalez, Y., Gil, D., de los Angeles Chavez, M., Betzel, C., Redecke, L.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 1289-1293
- PubMed: 23143234 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112039085
- Primary Citation Related Structures:
3OFW - PubMed Abstract:
The BPTI/Kunitz-type inhibitor family includes several extremely potent serine protease inhibitors. To date, the inhibitory mechanisms have only been studied for mammalian inhibitors. Here, the first crystal structure of a BPTI/Kunitz-type inhibitor from a marine invertebrate (rShPI-1A) is reported to 2.5 Å resolution. Crystallization of recombinant rShPI-1A required the salt-induced dissociation of a trypsin complex that was previously formed to avoid intrinsic inhibitor aggregates in solution. The rShPI-1A structure is similar to the NMR structure of the molecule purified from the natural source, but allowed the assignment of disulfide-bridge chiralities and the detection of an internal stabilizing water network. A structural comparison with other BPTI/Kunitz-type canonical inhibitors revealed unusual ϕ angles at positions 17 and 30 to be a particular characteristic of the family. A significant clustering of ϕ and ψ angle values in the glycine-rich remote fragment near the secondary binding loop was additionally identified, but its impact on the specificity of rShPI-1A and similar molecules requires further study.
- Centro de Estudio de Proteínas, Facultad de Biología, Universidad de la Habana, Calle 25 No. 455, Havana 10400, Cuba.
Organizational Affiliation: