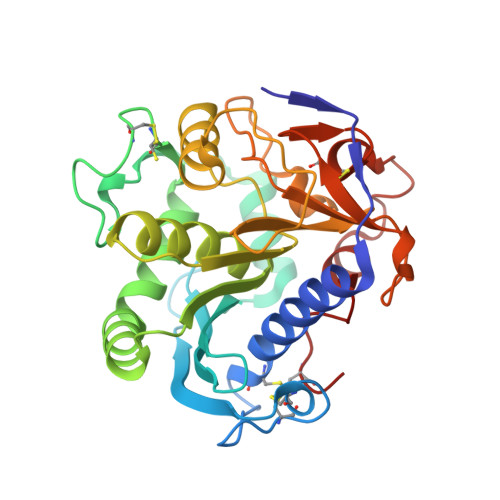
Exploring the conformational states and rearrangements of Yarrowia lipolytica Lipase.
Bordes, F., Barbe, S., Escalier, P., Mourey, L., Andre, I., Marty, A., Tranier, S.(2010) Biophys J 99: 2225-2234
- PubMed: 20923657 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2010.07.040
- Primary Citation Related Structures:
3O0D - PubMed Abstract:
We report the 1.7 Å resolution crystal structure of the Lip2 lipase from Yarrowia lipolytica in its closed conformation. The Lip2 structure is highly homologous to known structures of the fungal lipase family (Thermomyces lanuginosa, Rhizopus niveus, and Rhizomucor miehei lipases). However, it also presents some unique features that are described and discussed here in detail. Structural differences, in particular in the conformation adopted by the so-called lid subdomain, suggest that the opening mechanism of Lip2 may differ from that of other fungal lipases. Because the catalytic activity of lipases is strongly dependent on structural rearrangement of this mobile subdomain, we focused on elucidating the molecular mechanism of lid motion. Using the x-ray structure of Lip2, we carried out extensive molecular-dynamics simulations in explicit solvent environments (water and water/octane interface) to characterize the major structural rearrangements that the lid undergoes under the influence of solvent or upon substrate binding. Overall, our results suggest a two-step opening mechanism that gives rise first to a semi-open conformation upon adsorption of the protein at the water/organic solvent interface, followed by a further opening of the lid upon substrate binding.
- Université de Toulouse, Toulouse, France.
Organizational Affiliation: