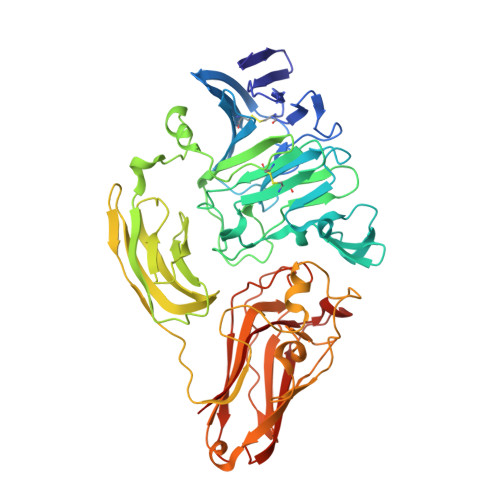
Structural and Biochemical Studies Elucidate the Mechanism of Rhamnogalacturonan Lyase from Aspergillus aculeatus.
Jensen, M.H., Otten, H., Christensen, U., Borchert, T.V., Christensen, L.L., Larsen, S., Leggio, L.L.(2010) J Mol Biology 404: 100-111
- PubMed: 20851126
- DOI: https://doi.org/10.1016/j.jmb.2010.09.013
- Primary Citation Related Structures:
2XHN, 3NJV, 3NJX - PubMed Abstract:
We present here the first experimental evidence for bound substrate in the active site of a rhamnogalacturonan lyase belonging to family 4 of polysaccharide lyases, Aspergillus aculeatus rhamnogalacturonan lyase (RGL4). RGL4 is involved in the degradation of rhamnogalacturonan-I, an important pectic plant cell wall polysaccharide. Based on the previously determined wild-type structure, enzyme variants RGL4_H210A and RGL4_K150A have been produced and characterized both kinetically and structurally, showing that His210 and Lys150 are key active-site residues. Crystals of the RGL4_K150A variant soaked with a rhamnogalacturonan digest gave a clear picture of substrate bound in the -3/+3 subsites. The crystallographic and kinetic studies on RGL4, and structural and sequence comparison to other enzymes in the same and other PL families, enable us to propose a detailed reaction mechanism for the β-elimination on [-,2)-α-l-rhamno-(1,4)-α-d-galacturonic acid-(1,-]. The mechanism differs significantly from the one established for pectate lyases, in which most often calcium ions are engaged in catalysis.
- Biophysical Chemistry Group, Department of Chemistry, University of Copenhagen, 2100 Copenhagen Ø, Denmark.
Organizational Affiliation: